At the König Lab, we merge the fields of Systems Medicine and Systems Biology to delve deep into drug
metabolism and liver function. By harnessing the power of computational modeling, data science,
bioinformatics analysis, and machine learning, we analyze and interpret datasets spanning biological,
medical, and clinical spheres.
A hallmark of our research is the generation of predictive digital twins mirroring individual human
physiology. These models are instrumental in shedding light on inter-individual variances, paving the way
for personalized treatments—from tailoring drug dosages to understanding the nuanced impacts of lifestyle on
drug metabolism. For more information, please see our
publications and projects.
We are actively involved in the standardization and reproducibility efforts in Systems Biology and Systems
Medicine.
We are actively involved in the standardization and reproducibility efforts in Systems Biology and Systems
Medicine.
For more information, see our
| 2026 |



|
TabularQual: A spreadsheet-based format for annotating and curating logical models in SBML-qual
Luna Xingyu Li, Carissa Bleker, Sylvain Soliman, Laurence Calzone, Noriko Hiroi, Tomas Helikar, Matthias König, Luiz Ladeira, Pedro Tiago Monteiro, Vincent Noel, Samuel Pastva, Albin Salazar, David Safranek, Denis Thieffry, Eirini Tsirvouli, Anna Niarakis, John Gennari
bioRxiv 2026.05.31.727710 (preprint). doi:10.64898/2026.05.31.727710
Logical models are widely used to study regulatory and signaling systems, yet their reuse, annotation, and exchange across tools remain challenging. Although SBML Level 3 Qualitative Models (SBML-qual) provides a standard representation, its XML-based syntax is difficult to inspect and edit directly. Here we introduce TabularQual, a spreadsheet-based, community-driven representation for Boolean and multi-valued logical models, together with a bidirectional converter between spreadsheets and SBML-qual. The converter is accessible programmatically and via a web interface to support diverse user workflows. We further describe two integration strategies that enable existing modeling tools to operate with TabularQual, either through SBML-qual exchange or via direct support. Case studies using the Stress Knowledge Map and CaSQ demonstrate how this integration supports model construction, curation, and reuse. TabularQual provides a practical bridge between human-readable model representations and standardized executable formats, supporting reproducibility, interoperability, and community-driven model development.
Keywords:
Logical models,
qual,
SBML,
standardization,
COMBINE,
|
|
| 2026 |


|
Reproducibility of a Physiologically Based Pharmacokinetic and Pharmacodynamic (PBPK/PD) Model of Dapagliflozin
Michelle Elias, Mariia Myshkina, Nike Nemitz, Matthias König
Physiome. May 13, 2026 (publication). doi:10.36903/physiome.31368556
A computational model in the form of a whole-body physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of dapagliflozin was developed to systematically evaluate the influence of patient-specific factors on drug disposition. Based on curated data from 28 clinical studies, the model simulates the absorption, distribution, metabolism and excretion (ADME) of the drug as well as its pharmacodynamics. The model accounts for variability in renal and hepatic function, and effects of food intake. The model is implemented in the Systems Biology Markup Language (SBML) standard. Analysis were performed utilizing the libroadrunner simulation library. Here, we demonstrate the computational reproducibility of the key findings from the primary publication, thereby verifying the consistency of the model implementation with the published results.
Keywords:
Dapagliflozin,
PBPK/PD,
SBML,
Pharmacokinetics,
Pharmacodynamics,
Computational Model,
|
|
| 2026 |


|
A PBPK Digital Twin of Rapamycin for Predicting Dose–Exposure Relationships, Organ Impairment Effects, and Drug–Drug Interactions
Monika Jesionek, Matthias König
Preprints 2026, 10.20944/preprints202605.0249.v1 (preprint). doi:10.20944/preprints202605.0328.v1
Sirolimus is a potent mTOR inhibitor used primarily to prevent acute rejection in transplant recipients. Its clinical management is challenging because of its narrow therapeutic index, low and highly variable oral bioavailability, and pronounced inter-individual variability driven by CYP3A4/5-mediated metabolism and P-glycoprotein efflux. Extensive partitioning into erythrocytes further complicates its disposition and necessitates therapeutic drug monitoring. Here, we developed a mechanistic whole-body physiologically based pharmacokinetic (PBPK) digital twin of rapamycin that integrates complex absorption kinetics, nonlinear distribution, and first-pass metabolism. The SBML-encoded model was calibrated and evaluated against a comprehensive library of curated clinical pharmacokinetic data, comprising studies primarily in healthy volunteers and stable renal transplant recipients. The dataset covers diverse ethnic populations, cohorts with varying degrees of renal and hepatic impairment, and individuals with relevant genetic polymorphisms. The digital twin captured overall trends in rapamycin blood concentrations across a wide range of doses and dosing regimens. Simulations showed good agreement with observed data under hepatic and renal impairment, as well as under fasted and fed conditions. Furthermore, the model reproduced the magnitude of drug--drug interactions involving potent CYP3A4 inhibitors, CYP3A4 inducers, and concomitant immunosuppressive agents. This SBML-based digital twin provides a quantitative framework for characterizing sirolimus dose dependency and the multifactorial effects of intrinsic and extrinsic factors on systemic exposure. By providing the model in a standards-based, executable format together with simulation scripts and curated pharmacokinetic datasets, this work supports independent reproduction, transparent model evaluation, and systematic reuse in accordance with FAIR principles.
Keywords:
rapamycin,
sirolimus,
PBPK modeling,
digital twin,
mTOR inhibitor,
drug–drug interactions,
organ impairment,
personalized medicine,
|
|
| 2026 |


|
An Open and Reproducible Digital Twin for Personalised Anticoagulant Therapy: Modelling Apixaban Pharmacokinetics and Pharmacodynamics
Mariia Myshkina, Mariia Babaeva, Linh Chang Nguyen, Shelee Bedón, Johannes Metternich, Michelle Elias, Matthias König
Preprints 2026, 10.20944/preprints202605.0249.v1 (preprint). doi:10.20944/preprints202605.0249.v1
Background: Thrombotic events increase with age, necessitating anticoagulants with reliable pharmacokinetic (PK) and pharmacodynamic (PD) profiles. Apixaban has important therapeutic advantages, but individualised dosing remains challenging because exposure and response are influenced by renal and hepatic function, food intake, body weight, and other patient-specific factors. Existing physiologically based pharmacokinetics/pharmacodynamics (PBPK/PD) models are limited by data sources, transparency, and incomplete representation of metabolites and pharmacodynamics. Methods: A systematic literature review identified 35 apixaban PK/PD clinical studies, which were curated and used for model development, parameter optimisation, and evaluation. We developed an expanded whole-body PBPK/PD model of apixaban with explicit metabolite representation and enhanced pharmacodynamic components. The model follows a modular structure and is encoded in SBML to support interoperability and reproducibility. Results: The model reproduced observed clinical PK/PD data across all 35 studies, covering diverse doses, regimens, and populations. Simulations captured apixaban PK and PD under normal conditions and clinically relevant scenarios, including renal and hepatic impairment, fasted and fed states, and obesity. Conclusions: This open PBPK/PD digital twin provides quantitative insight into determinants of apixaban exposure and response. All model files, documentation, simulation scripts, and curated datasets are openly available under MIT and CC-BY licenses following FAIR principles.
Keywords:
apixaban; anticoagulation,
pharmacokinetics/pharmacodynamics,
PBPK/PD modelling,
digital twin,
personalised medicine,
special populations,
FAIR data,
reproducible modelling,
|
|
| 2026 |


|
Automated Segmentation of Hepatic Vessels and Lobules in Whole-Slide Images Using U-Net Models
Mehul Bafna, Matthias König, Sylvia Saalfeld, Vladimira Moulisova, Vaclav Liska, Uta Dahmen, Mohamed Albadry
Front. Bioinform. 6:1713736 (publication). doi:10.3389/fbinf.2026.1713736
Automated analysis of hepatic vascular structures and lobules within whole-slide histological images is critical for ensuring accurate and timely morphometric evaluations and facilitating advancements in computational liver histology. Nonetheless, the intricate morphology of the tissue, variability in staining techniques, and the requirements for high-resolution imaging present substantial challenges to the precision of segmentation processes. We present a robust deep-learning pipeline using adaptive patch extraction and specialized U-Net architectures for segmenting vessels, bile ducts, and lobules in Glutamine Synthetase and Picro-Sirius-Red stained porcine liver sections. Our architecture incorporates a weight-boosted nnU-Net framework to effectively manage class imbalances and improve the detection of smaller vascular structures. Geometric data transformations enhanced the robustness and generalizability of the segmentation models. Evaluations conducted through five-fold cross-validation, as well as assessments utilizing independent test datasets, resulted in Dice similarity scores: 0.960 for lobules, 0.801 for central veins, 0.909 for hepatic arteries, 0.609 for portal veins, and 0.710 for bile ducts. The developed segmentation pipeline additionally supports comprehensive morphometric analyses of structural parameters, including number and size (diameter, area) of vascular structures, bile ducts, and lobules. For e.g., the diameter of hepatic arteries ranges between 30-90 µm. These findings underscore the practical relevance of adaptable segmentation frameworks in advancing computational histological analysis of liver tissue.
Keywords:
Hepatic Vascular Segmentation,
Lobule Detection,
Histological Image,
Analysis,
Deep Learning,
Lobule Segmentation,
|
|
| 2026 |


|
Parameter Uncertainty in the Optimization of Pharmacokinetic Models: A Reproducible Bayesian Approach
Antonio Alavarez (supervisor: Matthias König)
Master Thesis, April 2026 (thesis)
Bayesian methods offer a principled framework for parameter estimation and uncertainty quantification in physiologically-based pharmacokinetic (PBPK) modeling, yet the sensitivity of these approaches to key workflow hyperparameters remains insufficiently characterized. This thesis presents a reproducible methodology for evaluating Bayesian Markov Chain Monte Carlo (MCMC) parameter estimation across three pharmacokinetic models of increasing complexity: a simple chain model, a basic pharmacokinetic model, and a clinically validated PBPK model of indocyanine green (ICG). Synthetic pharmacokinetic datasets were generated from known parameter distributions and used to fit Bayesian MCMC samplers under systematically varied hyperparameters — number of timepoints, number of samples, coefficient of variation, and prior type. Posterior accuracy, point bias, computational cost, and sampling efficiency were evaluated across all conditions. Among the four hyperparameters investigated, prior specification emerged as the dominant determinant of posterior accuracy: well-specified priors yielded parameter estimates close to the true values across all models, while biased and incorrect priors introduced substantial and progressive deviations. The remaining hyperparameters had limited influence on estimation accuracy but the number of timepoints and samples directly increased computational cost. The entire workflow — built on SBML, libRoadRunner, PEtab, and pyPESTO — is fully open-source and FAIR-compliant, providing a transparent and reusable framework for validating Bayesian approaches in pharmacokinetic modeling.
Keywords:
Baysian,
parameter optimization,
SBML,
PEtab,
|
|
| 2026 |

|
From FAIR to CURE: Guidelines for Computational Models of Biological Systems
Herbert M. Sauro, Eran Agmon, Michael L. Blinov, John H. Gennari, Joe Hellerstein, Adel Heydarabadipour, Peter Hunter, Bartholomew E. Jardine, Elebeoba May, David P. Nickerson, Lucian P. Smith, Gary D Bader, Frank Bergmann, Patrick M. Boyle, Andreas Drager, James R. Faeder, Song Feng, Juliana Freire, Fabian Frohlich, James A. Glazier, Thomas E. Gorochowski, Tomas Helikar, Stefan Hoops, Princess Imoukhuede, Sarah M. Keating, Matthias König, Reinhard Laubenbacher, Leslie M. Loew, Carlos F. Lopez, William W. Lytton, Andrew McCulloch, Pedro Mendes, Chris J. Myers, Jerry G. Myers, Lealem Mulugeta, Anna Niarakis, David D. van Niekerk, Brett G. Olivier, Alexander A. Patrie, Ellen M. Quardokus, Nicole Radde, Johann M. Rohwer, Sven Sahle, James C. Schaff, T.J. Sego, Janis Shin, Jacky L. Snoep, Rajanikanth Vadigepalli, H. Steve Wiley, Dagmar Waltemath, Ion Moraru
npj Syst Biol Appl (2026) (publication). doi:10.1038/s41540-026-00651-0
Guidelines for managing scientific data have been established under the FAIR principles requiring that data be Findable, Accessible, Interoperable, and Reusable. In many scientific disciplines, especially computational biology, both data and models are key to progress. For this reason, and recognizing that such models are a very special type of 'data', we argue that computational models, especially mechanistic models prevalent in medicine, physiology and systems biology, deserve a complementary set of guidelines. We propose the CURE principles, emphasizing that models should be Credible, Understandable, Reproducible, and Extensible. We delve into each principle, discussing verification, validation, and uncertainty quantification for model credibility; the clarity of model descriptions and annotations for understandability; adherence to standards and open science practices for reproducibility; and the use of open standards and modular code for extensibility and reuse. We outline recommended and baseline requirements for each aspect of CURE, aiming to enhance the impact and trustworthiness of computational models, particularly in biomedical applications where credibility is paramount. Our perspective underscores the need for a more disciplined approach to modeling, aligning with emerging trends such as Digital Twins and emphasizing the importance of data and modeling standards for interoperability and reuse. Finally, we emphasize that given the non-trivial effort required to implement the guidelines, the community moves to automate as many of the guidelines as possible.
Keywords:
FAIR,
CURE,
reproducibility,
|
|
| 2026 |


|
A Digital Twin of Empagliflozin Pharmacokinetics and Pharmacodynamics
Jennesse Alejandro, Michelle Elias, Mariia Babaeva, Matthias König
Preprints 2026, 10.20944/preprints202603.1559.v1 (preprint). doi:10.20944/preprints202603.1559.v1
Background/Objectives: Empagliflozin is an SGLT2 inhibitor prescribed for the management of type 2 diabetes mellitus, lowering blood glucose by increasing urinary glucose excretion (UGE) through inhibition of renal glucose reabsorption. PK/PD responses vary substantially across patient populations, complicating dose selection under altered organ function. Here, we developed a whole-body PBPK/PD digital twin integrating absorption, distribution, metabolism, and excretion with explicit modeling of renal glucose handling via the renal threshold for glucose. Methods: The model represents empagliflozin and its glucuronide metabolite, is implemented in SBML, and was calibrated and evaluated against curated PK/PD data from 27 clinical studies spanning healthy individuals, patients with type 2 diabetes, and cohorts with renal or hepatic impairment. Results: The model accurately captured observed clinical PK/PD data across all 27 studies, spanning a wide range of doses, dosing regimens, and patient populations. Good agreement between simulations and observations was obtained under normal and impaired renal and hepatic function, as well as under fasted and fed conditions, demonstrating the model's ability to reproduce empagliflozin disposition and pharmacodynamic response across clinically relevant scenarios. Conclusions: This SBML-based PBPK/PD digital twin provides quantitative insight into empagliflozin dose dependency and the impact of renal impairment, hepatic impairment, and food intake on PK/PD across clinically relevant populations. All model files, simulation scripts, and curated datasets are openly available in accordance with FAIR principles.
Keywords:
digital twin,
empagliflozin,
Type 2 diabetes mellitus,
physiologically based pharmacokinetic/pharmacodynamic model (PBPK/PD),
pharmacokinetics,
pharmacodynamics,
personalized medicine,
|
|
| 2026 |



|
User-driven development and evaluation of an agentic framework for analysis of large pathway diagrams
Marie Corradi, Ivo Djidrovski, Luiz Ladeira, Bernard Staumont, Anouk Verhoeven, Julen Sanz Serrano, Adrien Rougny, Ahmad Vaez, Ahmed Hemedan, Alexander Mazein, Anna Niarakis, Arthur de Carvalho e Silva, Charles Auffray, Egon Wilighagen, Eliska Kuchovska, Falk Schreiber, Irina Balaur17, Laurence Calzone, Lisa Matthews, Lorenzo Veschini, Marc E Gillespie, Martina Kutmon, Matthias König, Matti van Welzen, Noriko Hiroi, Oxana Lopata, Pierre Klemmer, Rupert Overall, Tim Hofer, Venkata Satagopam, Reinhard Schneider, Marc Teunis, Liesbet Geris, Marek Ostaszewski
bioRxiv 2026.03.10.710813 (preprint). doi:10.64898/2026.03.10.710813
As biomedical knowledge keeps growing, resources storing available information multiply and grow in size and complexity. Such resources can be in the format of molecular interaction maps, which represent cellular and molecular processes under normal or pathological conditions. However, these maps can be complex and hard to navigate, especially to novice users. Large Language Models (LLMs), particularly in the form of agentic frameworks, have emerged as a promising technology to support this exploration. In this article, we describe a user-driven process of prototyping, development, and user testing of Llemy, an LLM-based system for exploring these molecular interaction maps. By involving domain experts from the very first prototyping in the form of a hackathon and collecting both fine-grained and general feedback on more refined versions, we were able to evaluate the perceived utility and quality of the developed system, in particular for summarising maps and pathways, as well as prioritise the development of future features. We recommend continued user-driven development and benchmarking to keep the community engaged. This will also facilitate the transition towards open-weight LLMs to support the needs of the open research environment in an ever-changing technology landscape.
Keywords:
SBGN,
LLM,
|
|
| 2026 |



|
SBML Level 3 Package: Flux Balance Constraints version 3
Brett G. Olivier, Frank T. Bergmann, Sarah Keating, and Matthias König
J Integr Bioinform. 2026 (accepted) (publication)
Constraint-based modeling is a well-established modeling methodology used to study biological networks at both the medium-scale and genome-scale. Due to their large size and complexity, such steady-state flux models are typically ana- lyzed using constraint-based optimization techniques, such as Flux Balance Anal- ysis (FBA). The Flux Balance Constraints (FBC) Package extends SBML Level 3 to provide a standardized format for encoding, exchanging, and annotating constraint-based models. It includes support for modeling concepts such as objective functions, flux bounds, and annotation of model components that facilitate reaction balanc- ing. Version two extended the original release by adding support for encoding gene-protein associations. Version three builds upon and maintains backwards compatibility with Version two by introducing new elements and attributes that include: user-defined constraints, a new quadratic variable type and key-value pair annotation that can be used to store additional information relevant to constraint-based modeling. The only changes to existing attributes are that a Species charge is no longer restricted to an integer value and chemicalFormula can now include generic components. In addition to providing the elements necessary to uniquely encode constraint- based models, the FBC package provides an open platform that facilitates the continued, cross-community development of an interoperable, constraint-based model encoding format.
Keywords:
SBML,
FBC,
|
|
| 2026 |



|
The Parameter Estimation tables (PEtab) format version 2
Dilan Pathirana, Fabian Fröhlich, Sebastian Persson, Frank T. Bergmann, Matthias König, Paul F. Lang, Severin Bang, Svenja Kemmer, Jan Hasenauer, and Daniel Weindl
J Integr Bioinform. 2026 (accepted) (publication)
Parameter estimation is a central task in data-driven computational modeling, and a clear, unambiguous, and commonly understood specification of parameter estimation problems is essential for reproducibility and interoperability. PEtab provides a standardized framework for defining such problems in systems biology and systems medicine, particularly for ordinary differential equation models. This document describes the second version of the PEtab format, PEtab 2.0. PEtab 2.0 simplifies and clarifies how estimation problems are defined, while also extending the standard to support more complex and realistic experimental setups.
Keywords:
Parameter estimation,
PETab,
|
|
| 2026 |


|
A Physiologically Based Pharmacokinetic and Pharmacodynamic (PBPK/PD) Model of Dapagliflozin in Type 2 Diabetes Mellitus: The Effect of Dosing, Hepatorenal Impairment, and Food
Nike Nemitz, Michelle Elias, Matthias König
Pharmaceutics 2026, 18(3), 287; https://doi.org/10.3390/pharmaceutics18030287 (publication). doi:10.3390/pharmaceutics18030287
Background/Objectives: Dapagliflozin is an SGLT2 inhibitor prescribed for the management of type 2 diabetes mellitus. The drug lowers blood glucose levels by increasing urinary glucose excretion (UGE). Despite established efficacy, dapagliflozin demonstrates significant inter-individual variability in pharmacokinetics (PK) and pharmacodynamics (PD), with potential impact on treatment outcomes. Methods: To evaluate the sources of variability and to support patient stratification and model-informed individualized therapy, we developed a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of dapagliflozin using curated data from 28 clinical studies. This framework integrates absorption, distribution, metabolism, excretion, and pharmacodynamics, and accounts for key determinants of variability including renal and hepatic function, and food effects. Results: The simulations reproduced dose-dependent pharmacokinetics with predicted Cmax and AUC values typically within 10–15% of observed data. Renal impairment reduced UGE by 40–60% despite modest changes in plasma exposure, while hepatic impairment produced only small shifts in PK and PD. The model also reproduced the fed-state reduction of peak concentrations, consistent with the 30–50% decrease reported clinically. Conclusions: All model files, code, and curated datasets are openly available in line with FAIR standards and Open Science practices, enabling transparent and reproducible analyses and providing a mechanistic basis for individualized therapy in type 2 diabetes.
Keywords:
dapagliflozin,
type 2 diabetes mellitus,
physiologically based pharmacokinetic/pharmacodynamic model (PBPK/PD),
pharmacokinetics,
pharmacodynamics,
personalized medicine,
|
|
| 2026 |


|
A Digital Twin of the Angiotensin II Receptor Blocker Losartan: Physiologically Based Modeling of Blood Pressure Regulation
Ennie Tensil, Mariia Myshkina, Matthias König
Pharmaceutics 2026, 18(2), 262; https://doi.org/10.3390/pharmaceutics18020262 (publication). doi:10.3390/pharmaceutics18020262
Background/Objectives: Losartan, an angiotensin II receptor blocker (ARB) used to treat hypertension and heart failure, shows significant variability in pharmacokinetics (PK) and pharmacodynamics (PD) among individuals. Methods: In this study, we developed a physiologically based pharmacokinetic/ pharmacodynamic (PBPK/PD) model of losartan and its active metabolite, E3174, using curated data from 25 clinical trials. The model mechanistically describes the processes of absorption, hepatic metabolism, renal and fecal excretion, and pharmacodynamic blood pressure regulation. Simulation studies examined the effects of dose, hepatic and renal impairment, and genetic polymorphisms in cytochrome p450 2C9 (CYP2C9) and P-glycoprotein 1, also known as multidrug resistance protein 1 (MDR1) or ATP-binding cassette sub-family B member 1 (ABCB1), on the model. Results: The model successfully reproduced key PK/PD observations, including dose-dependent receptor blockade, attenuated responses with hepatic impairment, modest enhancement with renal impairment, and substantial variability in E3174 formation dependent on CYP2C9; the effects of ABCB1 were minimal. Specifically, dose dependency simulations confirmed the saturable nature of CYP2C9 metabolism, predicting a decreasing E3174-to-losartan ratio and a stronger, sustained suppression of blood pressure and aldosterone at higher doses. Hepatic impairment was predicted to lead to elevated losartan plasma concentrations (increased AUC) and attenuated metabolite formation, confirming the clinical need for dose reduction. Renal impairment simulations predicted stable losartan AUC but showed an overestimation of E3174 accumulation compared to observed data, where E3174 exposure remained stable. Genetic variability (CYP2C9) was the major determinant of response, with simulations confirming that reduced-function alleles lead to a 1.6- to 3-fold increase in losartan AUC and diminished blood pressure reduction. ABCB1 variability resulted in only minor modulation of systemic exposure and blood pressure effects. Conclusions: This mechanistic digital twin framework provides a quantitative basis for understanding variability in losartan therapy and supports its application in individualized dosing strategies.
Keywords:
losartan,
angiotensin II receptor blocker,
ARB,
PBPK/PD modeling,
pharmacokinetics,
pharmacodynamics,
SBML,
|
|
| 2026 |


|
Reproducibility of a Digital Twin of the Angiotensin II Receptor Blocker Losartan
Mariia Myshkina, Michelle Elias, Ennie Tensil, Matthias König
Physiome. February 2, 2026 (publication). doi:10.36903/physiome.31359823
A digital twin in the form of a whole-body physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of losartan was developed to systematically evaluate the influence of patient-specific factors on drug disposition and effects. Based on curated data from 25 clinical studies, the model simulates the absorption, distribution, metabolism and excretion (ADME) as well as pharmacological effects of the drug. The model accounts for variability caused by the differences in renal and hepatic function, and by genetic polymorphisms of CYP2C9 and ABCB1. The model is implemented in the Systems Biology Markup Language (SBML) standard. Simulations were performed utilising the libroadrunner library. Here, we demonstrate the computational reproducibility of the key findings from the primary publication, thereby verifying the consistency and reproducibility of the model implementation with the published results.
Keywords:
Losartan,
PBPK/PD,
SBML,
Pharmacokinetics,
Pharmacodynamics,
Computational Model,
|
|
| 2026 |


|
A Digital Twin of Canagliflozin Pharmacokinetics and Pharmacodynamics in Type 2 Diabetes Mellitus
Vera Tereshchuk, Michelle Elias, Matthias König
Preprints 2026, 10.20944/preprints202601.2095.v1 (preprint). doi:10.20944/preprints202601.2095.v1
Canagliflozin is an SGLT2 inhibitor prescribed for the management of type 2 diabetes mellitus, lowering blood glucose by increasing urinary glucose excretion (UGE). However, pharmacokinetic (PK) and pharmacodynamic (PD) responses vary across patient populations, complicating dose selection under altered organ function. Here, we developed a whole-body PBPK/PD digital twin of canagliflozin that integrates absorption, distribution, metabolism, and excretion, and explicitly models renal glucose handling. The model represents canagliflozin and its major metabolites (M5, M7, M9) and was calibrated and evaluated using curated PK/PD data from 22 clinical studies spanning healthy individuals, patients with type 2 diabetes, and cohorts with renal or hepatic impairment. Simulations reproduced dose-dependent exposure and UGE across single- and multiple-dose regimens. Renal impairment produced modest changes in parent drug exposure but markedly reduced UGE and increased metabolite exposure, consistent with reduced glucose filtration and impaired metabolite clearance. Under hepatic impairment, simulations predicted increased canagliflozin exposure with altered metabolite profiles, while PD effects were minimal, although evaluation was limited by sparse clinical PD endpoints. All model files, simulation scripts, and curated datasets are provided in open SBML workflows in accordance with FAIR principles, enabling reproducible simulations and reuse for model-informed analyses of canagliflozin PK/PD variability.
Keywords:
Digital twin,
Canagliflozin,
Type 2 diabetes mellitus,
Physiologically based pharmacokinetic/pharmacodynamic model (PBPK/PD),
Pharmacokinetics,
Pharmacodynamics,
Personalized medicine,
|
|
| 2025 |


|
A Digital Twin of Glimepiride for Personalized and Stratified Diabetes Treatment
Michelle Elias, Matthias König
Front. Pharmacol. 16:1686415. (publication). doi:10.3389/fphar.2025.1686415. pmid:41132528
Introduction: Optimizing glimepiride therapy for type 2 diabetes (T2DM) is challenged by pronounced inter-individual variability in pharmacokinetics. Methods: We developed a whole-body physiologically based pharmacokinetic (PBPK) model as a digital twin of glimepiride, enabling systematic evaluation of how patient-specific factors influence drug disposition. Using curated data from 20 clinical studies, the digital twin mechanistically simulates glimepiride’s absorption, distribution, metabolism, and excretion (ADME). It accounts for key determinants of variability including renal and hepatic function, CYP2C9 genotype, and bodyweight. Results: The model accurately reproduced observed pharmacokinetics and quantified these factors’ impact on drug exposure. Increased glimepiride exposure was predicted in individuals with hepatic dysfunction or specific CYP2C9 variants, highlighting substantial genetic and physiological effects. Discussion: This digital twin provides mechanistic insights into pharmacokinetic variability and serves as an in silico platform for exploring individualized dosing and patient stratification strategies, laying the foundation for clinical decision support tools to improve T2DM management.
Keywords:
digital twin,
diabetes,
Glimepiride,
PBPK,
physiologically based pharmacokinetic model,
|
|
| 2025 |


|
Reproducibility of a Digital Twin of Glimepiride for Personalized and Stratified Diabetes Treatment
Michelle Elias, Matthias König
Physiome. October 20, 2025 (publication). doi:10.36903/physiome.28379193
A digital twin in the form of a whole-body physiologically based pharmacokinetic (PBPK) model of glimepiride was developed to systematically evaluate the influence of patient-specific factors on drug disposition. Based on curated data from 20 clinical studies, the model simulates the absorption, distribution, metabolism and excretion (ADME) of the drug while accounting for variability in renal and hepatic function, CYP2C9 genetic variants and bodyweight. The model is implemented in the Systems Biology Markup Language (SBML) standard and simulations are per- formed using scripts that utilise the libRoadRunner library to run simulations and generate results. Here, we demonstrate the computational reproducibility of the key findings from the primary publication, thereby verifying the consistency and reproducibility of the model implementation with the published results.
Keywords:
digital twin,
diabetes,
Glimepiride,
PBPK,
physiologically based pharmacokinetic model,
|
|
| 2025 |


|
Assessing the Impact of AI and Digital Twins on Clinical Decision-Making in Hepatology and Hepatobiliary Surgery
Mariia Myshkina, Elisabetta Casabianca, Anton Schnurpel, Tim Ricken, Hans-Michael Tautenhahn, Matthias König
Preprints 2025 (preprint). doi:10.20944/preprints202509.1164.v1
Background: Artificial Intelligence (AI) and Digital Twins (DT) are transforming clinical decision-making, particularly in hepatology and hepatobiliary surgery. While these technologies promise more precise diagnostics and individualised treatment planning, their successful integration depends on clinician acceptance, trust, and regulatory clarity. Methods: We conducted an online survey among hepatologists, hepatobiliary surgeons, and related specialists to assess familiarity, perceived value, adoption of clinical decision support systems (CDSS), concerns regarding automation bias and liability, and attitudes toward patient-centred integration. Data from 18 respondents were analysed using descriptive statistics and visualised in Python. Results: Most clinicians rated AI (80%) and DT (79%) as valuable to very valuable. Adoption of CDSS was high, with 93% reporting current or intended use. However, significant concerns emerged: 71% feared automation bias, and 92% expressed uncertainty about legal protection. Additionally, 77% supported incorporating patient preferences into AI- and DT-assisted decisions. Conclusions: Clinicians demonstrate a strong interest in adopting AI and DT, but remain cautious due to concerns over trust, liability, and regulatory gaps. Clear guidelines, clinical validation, and patient-centred approaches are essential for safe and effective integration into hepatology practice.
Keywords:
Digital Twins,
Artificial Intelligence,
Liver,
Hepatology,
Hepatobiliary Surgery,
Survey,
|
|
| 2025 |


|
A Systems Pharmacology Approach to Rivaroxaban: Physiologically Based Modeling of Pharmacokinetics and Coagulation Dynamics
Elisabetta Casabianca (supervisor: Matthias König)
Master Thesis, July 2025 (thesis)
Rivaroxaban, a direct oral anticoagulant (DOAC), is a widely used drug for the prevention and treatment of thromboembolic disorders. While its fixed dosing regimen and predictable anticoagulation profile improve clinical usability, significant inter-individual variability in pharmacokinetics (PK) and pharmacodynamics (PD) exists, particularly in patients with altered physiology or comorbidities. This study established an open database of rivaroxaban PK and PD data from 14 clinical studies. The data were used to develop and validate a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of rivaroxaban that captures its absorption, distribution, metabolism, excretion, and coagulation effects. The model incorporates the effects of dose, food intake, hepatic and renal function, and allows simulation of activated Factor X (Factor Xa) inhibition, prothrombin time (PT), and activated partial thromboplastin time (aPTT). Simulations accurately reproduced observed plasma concentration-time and coagulation profiles across a range of doses, fed/fasting conditions, as well as with different degrees of cirrhosis and renal impairment. This mechanistic PBPK/PD model provides a framework for exploring variability in rivaroxaban response across diverse clinical scenarios and supports its use in personalized dosing strategies, clinical trial design, and regulatory evaluation.
Keywords:
rivaroxaban,
direct oral anticoagulant,
DOAC,
PBPK/PD modeling,
pharmacokinetics,
pharmacodynamics,
|
|
| 2025 |


|
A Systems Pharmacology Approach to Rivaroxaban: Physiologically Based Modeling of Pharmacokinetics and Coagulation Dynamics
Elisabetta Casabianca, Mariia Myshkina, Matthias König
Preprints 2025, 202507.1945. (preprint). doi:10.20944/preprints202507.1945.v1
Rivaroxaban, a direct oral anticoagulant (DOAC), is a widely used drug for the prevention and treatment of thromboembolic disorders. While its fixed dosing regimen and predictable anticoagulation profile improve clinical usability, significant inter-individual variability in pharmacokinetics (PK) and pharmacodynamics (PD) exists, particularly in patients with altered physiology or comorbidities. This study established an open database of rivaroxaban PK and PD data from 14 clinical studies. The data were used to develop and validate a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of rivaroxaban that captures its absorption, distribution, metabolism, excretion, and coagulation effects. The model incorporates the effects of dose, food intake, hepatic and renal function, and allows simulation of activated Factor X (Factor Xa) inhibition, prothrombin time (PT), and activated partial thromboplastin time (aPTT). Simulations accurately reproduced observed plasma concentration-time and coagulation profiles across a range of doses, fed/fasting conditions, as well as with different degrees of cirrhosis and renal impairment. This mechanistic PBPK/PD model provides a framework for exploring variability in rivaroxaban response across diverse clinical scenarios and supports its use in personalized dosing strategies, clinical trial design, and regulatory evaluation.
Keywords:
rivaroxaban,
direct oral anticoagulant,
DOAC,
PBPK/PD modeling,
pharmacokinetics,
pharmacodynamics,
|
|
| 2025 |


|
A physiological-based pharmacokinetic/ pharmacodynamic (PBPK/PD) model of the angiotensin II receptor blocker (ARB) losartan
Ennie Tensil (supervisor: Matthias König)
Bachelor Thesis, July 2025 (thesis)
Losartan is an angiotensin II receptor antagonist commonly used in the treatment of hyperten- sion and heart failure. Its pharmacokinetics and pharmacodynamics are influenced by a range of physiological and genetic factors, which may affect therapeutic efficacy. This thesis presents a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of losartan and its active metabolite E3174 to explore these influences systematically. The modular SBML-based model includes submodels for absorption, hepatic metabolism, renal excretion, and pharmacodynamic effects via the renin-angiotensin-aldosterone system (RAAS). Simulations were conducted to investigate the impact of dose, hepatic and renal impairment, as well as CYP2C9 and ABCB1 polymorphisms. The model successfully reproduced observed pharmacological trends, including a dose-dependent increase in pharmacodynamic response, likely reflecting enhanced receptor blockade at higher systemic exposures. Hepatic dysfunction blunted the pharmacodynamic response to losartan, whereas renal impairment reduced metabolite clearance and lead to modest enhancement in pharmacodynamic effect. Genetic variability in CYP2C9 significantly altered E3174 formation and downstream RAAS inhibition, whereas ABCB1 activity had only minor effects on systemic exposure. The model provides mechanistic insight into inter-individual variability in losartan therapy and supports its potential use in individualized dosing strategies.
Keywords:
PK/PD,
pharmacokinetics,
pharmacodynamics,
losartan,
hepatorenal impairment,
pharmacogenomics,
food effect,
|
|
| 2025 |


|
Physiologically-Based Pharmacokinetic/Pharmacodynamic Modeling of Dapagliflozin: Exploring the Impact of Dosing, Hepatorenal Impairment and Food Intake
Nike Nemitz (supervisor: Matthias König)
Bachelor Thesis, June 2025 (thesis)
Dapagliflozin is a SGLT2 inhibitor used in the management of type 2 diabetes. Its effect is based on the reduction of plasma glucose levels by promoting glucose excretion in the urine. The pharmacokinetics and pharmacodynamics of dapagliflozin exhibit considerable interindividual variability, which is influenced by patient-specific factors and potentially affects therapeutic outcomes.
In this study, a physiologically based pharmacokinetic/ pharmacodynamic (PBPK/PD) model was developed to systematically assess how these factors influence dapagliflozin disposition. Using curated data from 27 clinical trials, a modular SBML-based model with sub-models for intestinal absorption, hepatic metabolism and renal excretion was developed.
Parameter optimization resulted in good agreement between simulations and observed clinical PK/PD profiles. The validated model accurately reproduced dose-dependent kinetics within the therapeutic range, captured the effects of physiological variability and functional impairments, and simulated food-drug interactions.
In addition, the model enabled direct comparison of urinary glucose excretion (UGE) under renal and hepatic impairment. Results showed that renal dysfunction substantially reduces UGE, while hepatic impairment has only a minor effect. Furthermore, the simulations revealed that food intake, although not altering total systemic exposure (AUC), significantly affects peak concentrations (Cmax), which in turn influences pharmacodynamic outcomes such as UGE. This suggests a clinically relevant food effect that is often underestimated.
Despite some limitations, the model offers mechanistic insight into the sources of PK/PD variability and supports individualized dosing strategies, contributing to predictive tools for personalized diabetes management and pharmacotherapy optimization.
Keywords:
PK/PD,
pharmacokinetics,
pharmacodynamics,
dapagliflozin,
hepatorenal impairment,
pharmacogenomics,
food effect,
ODE,
|
|
| 2025 |


|
A physiological-based pharmacokinetic (PBPK) model of the sulfonylurea glimepiride
Michelle Elias (supervisor: Matthias König)
Bachelor Thesis, April 2025 (thesis)
Glimepiride is a second-generation sulfonylurea used in the management of type 2 diabetes. Its pharmacokinetics exhibit significant inter-individual variability influenced by patient-specific factors, potentially affecting therapeutic outcomes. This study developed a physiologically based pharmacokinetic (PBPK) model to systematically evaluate how these factors affect glimepiride disposition. Using curated data from 19 clinical studies, a modular SBML-based model was developed with submodels for intestinal absorption, hepatic metabolism, and renal excretion. Parameter optimization achieved good alignment with observed pharmacokinetic profiles. The validated model reproduced dose-proportional kinetics within the therapeutic range and pre- dicted differential impacts of physiological variability. Renal impairment primarily affected metabolite clearance, while hepatic dysfunction led to increased glimepiride exposure via reduced cytochrome P450 2C9 (CYP2C9)-mediated metabolism. Bodyweight and CYP2C9 variants also influenced drug disposition. Despite CYP2C9 genotypes significantly affected individual pharmacokinetics, population-level differences across biogeographical groups were modest. While some limitations remain, particularly regarding data availability in severe organ dysfunction, the model offers mechanistic insights into pharmacokinetic variability. It provides a basis for individualized dosing strategies and supports the ongoing development of clinically applicable decision support tools for type 2 diabetes therapy.
Keywords:
PK/PD,
glimepiride,
hepatorenal impairment,
pharmacogenomics,
ODE,
CYP2C9,
|
|
| 2025 |

|
Inflammation and autophagy in peripheral nerves of rodent models with metabolic syndrome and type 2 diabetes mellitus
Baum P, Ebert T, Klöting N, Krupka S, König M, Paeschke S, Stock P, Bulc M, Blüher M, Palus K, Nowicki M, Kosacka J.
Neurosci Res. 2025 Apr 17:S0168-0102(25)00070-7 (publication). doi:10.1016/j.neures.2025.04.002. pmid:40252698
Metabolic syndrome (MetS) and type 2 diabetes mellitus (T2D) are associated with inflammation and the accumulation of macrophages in peripheral nerves, which increases the risk of developing peripheral neuropathy (PN). We have previously investigated that macrophage infiltration in the peripheral nerves of animals with T2D (leptin-deficient ob/ob mice, leptin receptor-deficient db/db) correlated with PN, whereas this process in animals with MetS (Wistar Ottawa Karlsburg W (RT1u) WOKW rat) did not lead to neuropathic changes. Additional data presented in this study suggest an association between increased mRNA expression of the anti-inflammatory marker IL-10 and autophagy in the prevention of neuropathy.
Keywords:
Autophagy,
Inflammation,
Metabolic syndrome;,
PDN,
T2D,
WOKW rat,
db/db mice,
ob/ob mice,
|
|
| 2025 |



|
FAIRification of computational models in biology
Irina Balaur, David P. Nickerson, Danielle Welter, Judith A.H. Wodke, Francois Ancien, Tom Gebhardt, Valentin Grouès, Henning Hermjakob, Matthias König, Nicole Radde, Adrien Rougny, Reinhard Schneider, Rahuman S. Malik-Sheriff, Kirubel Biruk Shiferaw, Melanie Stefan, Venkata Satagopam, Dagmar Waltemath
bioRxiv 2025.03.21.644517 (preprint). doi:10.1101/2025.03.21.644517
Computational models are essential for studying complex systems which, particularly in clinical settings, need to be quality-approved and transparent. To enhance the communication of a model’s features and capabilities, we propose an adaptation of the Findability, Accessibility, Interoperability and Reusability (FAIR) indicators published by the Research Data Alliance to assess models encoded in domain-specific standards, such as those established by COMBINE. The assessments guide FAIRification and add value to models.
Keywords:
FAIR,
computational models,
|
|
| 2025 |


|
A Physiologically-Based Pharmacokinetic Model of Tirzepatide
Internship Report Abhinav Mishra (supervisor: Matthias König)
Internship Report, Mar 2025 (report)
Tirzepatide, a dual glucose-dependent insulinotropic polypeptide (GIP) and glucagon-like peptide-1 (GLP-1) receptor agonist, represents a novel approach for treating type 2 diabetes mellitus (T2DM) and obesity. By combining the actions of both incretins, it enhances insulin secretion, suppresses glucagon, and reduces appetite, leading to superior glycemic control and weight loss compared to traditional GLP-1 receptor agonists. Despite its efficacy, inter-individual variability in pharmacokinetics necessitates a mechanistic understanding for optimized therapeutic application. This study employs physiologically-based pharmacokinetic (PBPK) modeling to characterize tirzepatide’s absorption, distribution, metabolism, and excretion (ADME). The model integrates physiological parameters to assess dose proportionality, hepatic and renal impairment effects, and body weight dependency. By improving drug exposure predictions, this approach aids personalized treatment strategies and enhances clinical decision-making across diverse patient populations.
Keywords:
PKDB,
tirzepatide,
GLP-1,
Pharmacokinetics,
|
|
| 2025 |

|
Anti-Endoglin monoclonal antibody prevents the progression of liver sinusoidal endothelial inflammation and fibrosis in MASH
Eissazadeh S, Fikrova P, Rathouska JU, Nemeckova I, Tripska K, Vasinova M, Havelek R, Mohammadi S, Igreja Sa IC, Theuer C, König M, Micuda S, Nachtigal P
Life Sci. 2025 Jan 29:123428 (publication). doi:10.1016/j.lfs.2025.123428. pmid:39889923
Liver sinusoidal endothelial inflammation/dysfunction and fibrosis are a crucial part of Metabolic Dysfunction Associated Steatohepatitis (MASH) development. TRC105 and M1043 are anti-endoglin (ENG) monoclonal antibodies that bind ENG. In this study, we hypothesized that treatment with anti-ENG antibodies would prevent the progression of LSECs inflammation and fibrosis in vivo and in vitro. MASH was induced in male C57BL/6 mice fed a choline-deficient L-amino acid-defined high-fat diet (CDAA-HFD) for 4 or 8 weeks. In the rescue study, mice were divided into three groups: a control group (chow diet), a MASH group (CDAA-HFD + IgG), and a rescue group (CDAA-HFD + M1043). Later, two groups received rat IgG1 (10 mg/kg) and M1043 (10 mg/kg). In in vitro experiments, inflammation was induced in human LSECs by ox-LDL (50 μg/mL) and treated with TRC105 (300 μg/mL). Liver sinusoidal endothelial inflammation/dysfunction in MASH animals was characterized by endothelial overexpression of ENG, VCAM-1, and ICAM-1 and reduced VE-cadherin and p-eNOS/eNOS expression. M1043 treatment prevented the overexpression of ENG, VCAM-1, and ICAM-1, the progression of liver fibrosis, and the increase of liver-to-body weight ratio. In vitro experiments with TRC105 confirmed the prevention of LSECs inflammation development by reduced ENG and VCAM-1 expression, as well as decreased THP-1 monocytic cell adhesion in ox-LDL activated LSECs. In conclusion, we demonstrate that anti-ENG antibody treatment can prevent LSECs inflammation and fibrosis progression in a MASH animal model and LSECs inflammation in vitro. Thus, we propose directly targeted ENG may represent a promising pharmacological approach for addressing LSECs inflammation and liver fibrosis.
Keywords:
Anti-endoglin antibody,
Endoglin,
Fibrosis,
Liver alteration,
Liver sinusoidal endothelial inflammation,
|
|
| 2025 |


|
Reproducible digital twins for personalized liver function assessment
Matthias König
Conference Proceedings, COMBINE 2024 - Conference of the Computational Modeling in Biology Network : September 1-5, 2024 (abstract). doi:10.24407/KXP:1902121317
Essential prerequisites for the practical application and translation of computational models include: i) reproducibility of results; ii) model reusability and extensibility; iii) data availability; and iv) strategies for model stratification and individualization. Here, we present a modeling workflow built around these foundational prerequisites, with a focus on liver function tests. Despite the paramount significance of liver function assessment in hepatology, reliable quantification remains a clinical challenge. Dynamic liver function tests offer a promising method for non-invasive in vivo assessment of liver function and metabolic phenotyping. By leveraging whole-body physiologically-based pharmacokinetic (PBPK) models, we are simulating these tests and positioning PBPK models as digital twins for metabolic phenotyping and liver function assessment. To develop and validate our models, we established the open pharmacokinetics database, PK-DB, containing curated data from 600+ clinical studies (10.1093/nar/gkaa990, 10.3389/fphar.2021.752826). Our models are individualizable and stratifiable, enabling simulation of lifestyle factors and co-administration effects on drug metabolism. Our models have been instrumental in clinical scenarios: from predicting individual outcomes post-hepatectomy (10.3389/fphys.2021.730418, 10.3389/fphys.2021.757293) to discerning the impact of CYP2D6 gene variants on liver function tests (10.3389/fphar.2022.1029073). These models are constructed hierarchically, describing metabolic and other biological processes in organs like the liver and kidneys, seamlessly integrated with whole-body physiology. Notably, all models and data are readily available and reproducible for reuse, encoded in the Systems Biology Markup Language (SBML) (10.15252/msb.20199110). We will provide an overview of these PBPK models and demonstrate how SBML and FAIR principles can facilitate model development, coupling, and reuse.
Keywords:
digital twin,
reproducibility,
liver,
FAIR,
pharmacokinetics,
|
|
| 2025 |


|
Computational Model Development Using SBML: sbmlutils, sbm4humans, cy3sbml
Matthias König
Conference Proceedings, COMBINE 2024 - Conference of the Computational Modeling in Biology Network : September 1-5, 2024 (abstract). doi:10.24407/KXP:1902121317
The Systems Biology Markup Language (SBML) (Keating et al., 2020) is recognized as the standard framework for representing and exchanging complex mathematical models in biological systems research. SBML facilitates the depiction of a diverse array of biological phenomena, encom- passing metabolic networks, signaling pathways, and regulatory networks. It is versatile enough to handle models ranging from simple individual processes to intricate multi-scale representations. One of the primary challenges faced by newcomers in computational biology is the encoding and development of ordinary differential equation (ODE) models within the SBML framework. Addressing this hurdle, we introduce two innovative Python tools: sbmlutils (https://github. com/matthiaskoenig/sbmlutils), sbml4humans (https://sbml4humans.de), and the Cytoscape application cy3sbml (https://github.com/matthiaskoenig/cy3sbml). These tools collectively streamline the process of SBML model creation, enhancing both the programmatic aspect and the user experience. Specifically, sbmlutils facilitates the programmatic construction of SBML models, while sbml4humans generates user-friendly reports for model interpretation. Furthermore, cy3sbml integrates with Cytoscape to offer advanced visualization capabilities, thereby augmenting the comprehension and analysis of SBML-encoded models. These advancements significantly contribute to the ease of SBML model development and interpretation, fostering greater accessibility and understanding for those entering the field of computational systems biology.
Keywords:
digital twin,
reproducibility,
liver,
|
|
| 2025 |


|
Physiologically based pharmacokinetic/pharmacodynamic modeling of the direct renin inhibitor aliskiren: Exploring the impact of hepatorenal impairment and drug-drug interactions
Bachelor Thesis Yusuf Ali Kulanoglu (supervisor: Matthias König)
Bachelor Thesis, January 2025 (thesis)
In this thesis, an extensive dataset of aliskiren pharmacokinetics and pharmacodynamics was compiled and used to develop a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model. This model was applied to address three primary research questions: I. How does hepatorenal impairment influence the pharmacokinetics of aliskiren?; II. What effects do co-administered medications have on the pharmacokinetics of aliskiren?; III. What are the consequences of these pharmacokinetic changes for the pharmacodynamics of aliskiren, particularly its ability to modulate the renin-angiotensin-aldosterone system (RAAS)?
The PBPK/PD model enables the prediction of aliskiren concentration-time profiles following oral and intravenous administration and provides detailed insights into the factors that affect its pharmacological efficacy. By exploring these influences, the study offers a foundation for developing optimized therapeutic strategies tailored to individual patients.
Keywords:
PK/PD,
aliskiren,
hepatorenal impairment,
ODE,
|
|
| 2024 |

|
FROG Analysis Ensures the Reproducibility of Genome Scale Metabolic Models
Karthik Raman, Miroslav Kratochvil, Brett G. Olivier, Matthias König, Pratyay Sengupta, Dinesh Kumar Kuppa Baskaran, Tung V N Nguyen, Daniel Lobo, St Elmo Wilken, Krishna Kumar Tiwari, Aswathy K. Raghu, Indumathi Palanikumar, Lavanya Raajaraam, Maziya Ibrahim, Sanjaay Balakrishnan, Shreyansh Umale, Frank Bergmann, Tanisha Malpani, Venkata P Satagopam, Reinhard Schneider, Moritz E. Beber, Sarah Keating, Mihail Anton, Alina Renz, Meiyappan Lakshmanan, Dong-Yup Lee, Lokanand Koduru, Reihaneh Mostolizadeh, Oscar Dias, Emanuel Cunha, Alexandre Oliveira, Yi Qing Lee, Karsten Zengler, Rodrigo Santibanez-Palominos, Manish Kumar, Matteo Barberis, Bhanwar Lal Puniya, Tomas Helikar, Hoang V. Dinh, Patrick F. Suthers, Costas D. Maranas, Isabella Casini, Seyed Babak Loghmani, Nadine Veith, Nantia Leonidou, Feiran Li, Yu Chen, Jens Nielsen, GaRyoung Lee, Sang Mi Lee, Gi Bae Kim, Pedro T. Monteiro, Miguel C. Teixeira, Hyun Uk Kim, Sang Yup Lee, Ulf W. Liebal, Lars M. Blank, Christian Lieven, Chaimaa Tarzi, Claudio Angione, Manga Enuh Blaise, Celik Pinar Aytar, Mikhail Kulyashov, llya Akberdin, Dohyeon Kim, Sung Ho Yoon, Zhaohui Xu, Jyotshana Gautam, William T. Scott Jr., Peter J. Schaap, Jasper J. Koehorst, Cristal Zuniga, Gabriela Canto-Encalada, Sara Benito-Vaquerizo, Ivette Parera Olm, Maria Suarez-Diez, Qianqian Yuan, Hongwu Ma, Mohammad Mazharul Islam, Jason A. Papin, Francisco Zorrilla, Kiran Raosaheb Patil, Arianna Basile, Juan Nogales, Granado San Leon, Freddy Castillo-Alfonso, Roberto Olivares-Hernandez, Gabriel Vigueras-Ramirez, Henning Hermjakob, Andreas Drager, Rahuman S Malik-Sheriff
bioRxiv 2024.09.24.614797 (preprint). doi:10.1101/2024.09.24.614797
Genome scale metabolic models (GEMs) and other constraint-based models (CBMs) play a pivotal role in understanding biological phenotypes and advancing research in areas like metabolic engineering, human disease modelling, drug discovery, and personalized medicine. Despite their growing application, a significant challenge remains in ensuring the reproducibility of GEMs, primarily due to inconsistent reporting and inadequate model documentation of model results. Addressing this gap, we introduce FROG analysis, a community driven initiative aimed at standardizing reproducibility assessments of CBMs and GEMs. The FROG framework encompasses four key analyses including Flux variability, Reaction deletion, Objective function, and Gene deletion to produce standardized, numerically reproducible FROG reports. These reports serve as reference datasets, enabling model evaluators, curators, and independent researchers to verify the reproducibility of GEMs systematically. BioModels, a leading repository of systems biology models, has integrated FROG analysis into its curation workflow, enhancing the reproducibility and reusability of submitted GEMs. In our study evaluating 65 GEM submissions from the community, approximately 40% reproduced without intervention, 28% requiring minor adjustments, and 32% needing input from authors. The standardization introduced by FROG analysis facilitated the detection and resolution of issues, ultimately leading to the successful reproduction of all models. By establishing a standardized and comprehensive approach to evaluating GEM reproducibility, FROG analysis significantly contributes to making CBMs and GEMs more transparent, reusable, and reliable for the broader scientific community.
Keywords:
COMBINE,
FROG,
FBC,
|
|
| 2024 |


|
Quantitative Image Analysis of Hepatic Zonation in Cytochrome P450 and Steatosis Using Whole Slide Scans
Master Thesis Jonas Küttner (supervisor: Matthias König)
Master Thesis, August 2024 (thesis)
The mammalian liver is organized into three-dimensional structures called lobules that are integral to its function. According to the widely accepted hexagonal model, the cross-section of each lobule is characterized by a central vein and portal triads at the corners. These portal triads consist of a hepatic portal vein, a hepatic arteriole, and a bile duct. Blood is delivered through the hepatic artery and portal vein and flows inward from the outer periportal region toward the central vein through sinusoids lined with hepatocytes. These hepatocytes express a wide variety of metabolic enzymes, including cytochrome P450 (CYP) isoforms that are critical for xenobiotic metabolism. Notably, many hepatic enzymes, including CYPs, exhibit differential expression along the periportal-to-perivenous axis, a spatial variation known as liver zonation.
Non-alcoholic fatty liver disease (NAFLD) is one of the most common liver diseases characterized by the accumulation of fat in the liver (steatosis) in the absence of significant alcohol consumption. Histopathologically, NAFLD is characterized by the presence of microvesicular fat droplets within hepatocytes.
Although liver zonation has been extensively studied, the protein gradients of CYPs and their alterations in response to steatosis have not been systematically quantified or compared across species. This study aimed to develop an image analysis workflow to accurately derive these lobular expression gradients and quantify macrosteatosis in whole slide images (WSI) of liver histology.
A comprehensive analysis was performed on whole slide images (WSIs) of mouse, rat, pig, and human liver specimens. These images included hematoxylin and eosin (H\&E) stained slides and immunohistochemically stained slides for glutamine synthetase (GS) and cytochrome P450 isoforms 1A2, 2E1, 2D6, and 3A4. To achieve the study objectives, two image analysis workflows were developed: one using classical methods and the other using convolutional neural networks (CNN). These workflows were designed to perform two key tasks: (1) segmentation of liver lobules and (2) detection of macrosteatosis droplets.
Key findings were that there was significant variability in lobular geometry within subjects, with strong consistency across species, with inter-species differences observed primarily in lobular size. A slight increase in lobular size was observed in steatotic samples, with a moderate to strong correlation between lobular size and steatosis content. In addition, interspecies differences in zonation patterns were observed for CYP1A2 in pigs, CYP2E1 in rats, and CYP3A4 in humans. Importantly, zonation patterns in steatotic subjects were not qualitatively different from those in non-steatotic subjects across species.
In summary, I have successfully developed and implemented an image analysis workflow for whole slide images (WSIs) that allows detailed examination of liver lobule geometry and macrosteatosis droplets. Furthermore, the resulting datasets were integrated to facilitate a comprehensive analysis of the zonal distribution of CYP protein expression and macrosteatosis within the lobules.
Keywords:
liver,
image analysis,
steatosis,
machine learning,
whole slide image (WSI),
|
|
| 2024 |

|
Modulation of endoglin and soluble endoglin in NASH: Computational modeling and monoclonal antibody therapeutic benefits
Samira Eissazadeh, Jana Urbankova Rathouska, Ivana Nemeckova, Matthias König, Petr Nachtigal
Atherosclerosis, Volume 395, Supplement 1, August 2024, 118102, EAS Abstracts 2024 (abstract). doi:10.1016/j.atherosclerosis.2024.118102
Background and Aims: NASH, a growing liver disease worldwide,requires an understanding of its molecular progression for effective treatment.Endoglin,a TGFβ-superfamily coreceptor,has two forms:membrane endoglin (ENG)and soluble endoglin (sENG).While ENG is associated with liver sinusoidal endothelial dysfunction (LSED)and liver fibrosis, its direct involvement in the progression of NASH remains unclear.Our aim was to investigate the relationship between NASH-progression,ENG expression,and LSED development.We hypothesized that the anti-endoglin monoclonal antibody M1043 would affect LSED development by directly affecting ENG expression and signaling with respect to NASH development.
Methods: NASH was induced in two separated groups of male C57BL/6 mice using the CDAA-HFD diet(n=7),while two control groups(n=7)were fed a standard diet.Mice were sacrificed after four and eight weeks to evaluate ENG protein changes during NASH. Then,24 mice were grouped into the control group(n=8),CDAA+rat IgG(n=8),and CDAA+M1043 group (n=8).After four weeks,CDAA+IgG and CDAA+M1043 groups received intraperitoneal injections of rat IgG (10 mg/kg)and M1043(10 mg/kg) twice a week.The experiment lasted eight weeks,with blood and liver samples collected for analysis.To better understand the role of ENG antibody and its cleavage,a computational model of ENG modulation of TGFβ signaling was developed.
Results: Liver injury was confirmed by elevated liver enzymes,increased fibrosis,and inflammatory biomarkers in both CDAA-HFD groups.LSEC development was confirmed by increased ENG,VCAM-1,and ICAM-1 expression in endothelial cells.We showed increased expression of MMP-14 and higher levels of sENG levels in 8 weeks fed mice.M1043 treatment significantly prevented the increase in ENG,VCAM-1,and ICAM-1 protein expression in liver sinusoidal endothelial cells when compared to non-treated mice.Computational modeling simulations showed the ENG-dependent switch between Smad2/3 and Smad1/5/8 signaling,along with changes in sENG levels,which was reversed by ENG antibodies.
Conclusions: Our findings indicate that NASH development is associated with LSED development,increased ENG and MMP-14 expression,and elevated sENG levels,suggesting ENG role in LSED progression.Blocking of ENG expression and signaling might be a potential target to affect LSED development and possibly prevent NASH aggravation.
Keywords:
endoglin,
modeling,
|
|
| 2024 |

|
Specifications of standards in systems and synthetic biology: status, developments, and tools in 2024
Martin Golebiewski, Gary Bader, Padraig Gleeson, Thomas E Gorochowski, Sarah M Keating, Matthias König, Chris J Myers, David P Nickerson, Björn Sommer, Dagmar Waltemath, Falk Schreiber
J Integr Bioinform. 2024 Jul 22 (publication). doi:10.1515/jib-2024-0015. pmid:39026464
The “COmputational Modeling in BIology NEtwork” (COMBINE) initiative aims to harmonise the development of diverse community standards for computational models in biology. It coordinates standard development to support the associated projects towards establishing a suite of compatible, interoperable and comprehensive standards that address the full spectrum of modeling in systems and synthetic biology.
Keywords:
COMBINE,
reproducibility,
standards,
|
|
| 2024 |

|
EFECT – A Method and Metric to Assess the Reproducibility of Stochastic Simulation Studies
T.J. Sego, Matthias König, Luis L. Fonseca, Baylor Fain, Adam C. Knapp, Krishna Tiwari, Henning Hermjakob, Herbert M. Sauro, James A. Glazier, Reinhard C. Laubenbacher, Rahuman S. Malik-Sheriff
arXiv:2406.16820 (preprint). doi:10.48550/arXiv.2406.16820
Reproducibility is a foundational standard for validating scientific claims in computational research. Stochastic computational models are employed across diverse fields such as systems biology, financial modelling and environmental sciences. Existing infrastructure and software tools support various aspects of reproducible model development, application, and dissemination, but do not adequately address independently reproducing simulation results that form the basis of scientific conclusions. To bridge this gap, we introduce the Empirical Characteristic Function Equality Convergence Test (EFECT), a data-driven, statistically robust method to quantify the reproducibility of stochastic simulation results. EFECT employs empirical characteristic functions to compare reported results with those independently generated by assessing distributional inequality, termed EFECT error, a metric to quantify the likelihood of equality. Additionally, we establish the EFECT convergence point, a quantitative metric for determining the required number of simulation runs to achieve an EFECT error value of a priori statistical significance, setting a reproducibility benchmark. EFECT supports all real-valued and bounded results irrespective of the model or method that produced them, and accommodates stochasticity from intrinsic model variability and random sampling of model inputs. We tested EFECT with stochastic differential equations, agent-based models, and Boolean networks, demonstrating its broad applicability and effectiveness. EFECT standardizes stochastic simulation reproducibility, establishing a workflow that guarantees reliable results, supporting a wide range of stakeholders, and thereby enhancing validation of stochastic simulation studies, across a model's lifecycle. To promote future standardization efforts, we are developing open source Stochastic Simulation Reproducibility software library (libSSR) in diverse programming languages for easy integration of EFECT.
Keywords:
COMBINE,
reproducibility,
|
|
| 2024 |


|
Cross-Species Variability in Lobular Geometry and Cytochrome P450 Hepatic Zonation: Insights into CYP1A2, CYP2E1, CYP2D6 and CYP3A4
Mohamed Albadry, Jonas Kuettner, Jan Grzegorzewski, Olaf Dirsch, Eva Kindler, Robert Klopfleisch, Vaclav Liska, Vladimira Moulisova, Sandra Nickel, Richard Palek, Jachym Rosendorf, Sylvia Saalfeld, Utz Settmacher, Hans-Michael Tautenhahn, Matthias KönigΔ, Uta DahmenΔ (Δ equal contribution)
Front Pharmacol. 2024 May 16;15:1404938 (publication). doi:10.3389/fphar.2024.1404938. pmid:38818378
There is a lack of systematic research exploring cross-species variation in liver lobular geometry and zonation patterns of critical drug-metabolizing enzymes, a knowledge gap essential for translational studies. This study investigated the critical interplay between lobular geometry and key cytochrome P450 (CYP) zonation in four species: mouse, rat, pig, and human. We developed an automated pipeline based on whole slide images (WSI) of hematoxylin-eosin-stained liver sections and immunohistochemistry. This pipeline allows accurate quantification of both lobular geometry and zonation patterns of essential CYP proteins. Our analysis of CYP zonal expression shows that all CYP enzymes (besides CYP2D6 with panlobular expression) were observed in the pericentral region in all species, but with distinct differences. Comparison of normalized gradient intensity shows a high similarity between mice and humans, followed by rats. Specifically, CYP1A2 was expressed throughout the pericentral region in mice and humans, whereas it was restricted to a narrow pericentral rim in rats and showed a panlobular pattern in pigs. Similarly, CYP3A4 is present in the pericentral region, but its extent varies considerably in rats and appears panlobular in pigs. CYP2D6 zonal expression consistently shows a panlobular pattern in all species, although the intensity varies. CYP2E1 zonal expression covered the entire pericentral region with extension into the midzone in all four species, suggesting its potential for further cross-species analysis. Analysis of lobular geometry revealed an increase in lobular size with increasing species size, whereas lobular compactness was similar. Based on our results, zonated CYP expression in mice is most similar to humans. Therefore, mice appear to be the most appropriate species for drug metabolism studies unless larger species are required for other purposes, e.g., surgical reasons. CYP selection should be based on species, with CYP2E1 and CYP2D6 being the most preferable to compare four species. CYP1A2 could be considered as an additional CYP for rodent versus human comparisons, and CYP3A4 for mouse/human comparisons. In conclusion, our image analysis pipeline together with suggestions for species and CYP selection can serve to improve future cross-species and translational drug metabolism studies.
Keywords:
liver,
zonation,
cytochrome P450,
lobule geometry,
drug metabolism,
species,
|
|
| 2024 |



|
Enhancing Our Understanding of Enalapril’s Pharmacokinetics: A Physiologically Based Modeling Approach
Master Thesis Shubhankar Palwankar (supervisor: Matthias König)
Master Thesis, May 2024 (thesis)
Enalapril is a medication used to treat high blood pressure and a num- ber of other cardiovascular conditions. The renin-angiotensin-aldosterone system (RAAS) is the main component involved in the regulation of blood pressure in the body. One of the key steps in this process is the conver- sion of angiotensin I to angiotensin II by angiotensin converting enzyme (ACE). Enalapril is an ACE inhibitor and therefore helps to lower blood pressure. Pharmacokinetics is the branch of pharmacology that studies how the body absorbs, distributes, metabolizes, and eliminates drugs over time. After administration as enalapril maleate, it is absorbed in the intestine and converted by carboxylesterase 1 (CES1) in the liver to the active metabolite, enalaprilat. Enalapril and enalaprilat are eliminated from the body by re- nal clearance. A physiologically based pharmacokinetic (PBPK) model was developed to investigate the pharmacokinetics of enalapril and the factors influencing it. As part of the work, an extensive database of enalapril phar- macokinetics consisting of data from 49 clinical trials was established and used to parameterize and validate the computational model. The model was used to investigate the effect of renal impairment, hepatic impairment and changes in CES1 activity on the pharmacokinetics of enalapril and enalapri- lat, as these three factors are known to have a major influence on the phar- macokinetics of enalapril. The model shows good agreement with a wide range of enalapril and enalaprilat data in serum, plasma and urine under different conditions (healthy, renal impairment, hepatic impairment, CES1 mutations). The model is available in SBML under a CC-BY 4.0 license with all data freely available from the PK-DB pharmacokinetics database.
Keywords:
hypertension,
enalapril,
PBPK,
pharmacokinetics,
CES1,
liver function,
renal function,
hepatic impairment,
renal impairment,
|
|
| 2024 |


|
A systematic overview of protein variability in cytochrome P450 and UDP-glucuronosyltransferase enzymes in the human liver
Bachelor Thesis Afruja Hossain (supervisor: Matthias König)
Bachelor Thesis, Jan 2024 (thesis)
Cytochrome P450 (CYP450) and UDP-glucuronosyltransferase (UGT) enzymes play key roles in the metabolism of a wide variety of endogenous and exogenous compounds, including drugs and toxins. CYP450 enzymes are involved in phase I metabolic reactions that typically introduce or unmask functional groups on molecules, often leading to their activation or inactivation. These reactions include hydroxylation, oxidation, and reduction. UGT enzymes, on the other hand, are essential for phase II metabolism, where they catalyze the conjugation of glucuronic acid to various substrates, making the compounds more water soluble and therefore easier to excrete. Both CYP450 and UGT are critical in determining drug pharmacokinetics, drug efficacy, drug interactions and potential toxicity.
A variety of CYP450 and UGT isoforms exist in the human liver, and different isoforms are responsible for detoxifying different substances. For example, CYP1A2 is involved in the metabolism of caffeine, CYP2D6 in dextromethorphan, CYP2E1 in chlorzoxazone, and CYP3A4 in midazolam. There are large inter-individual differences in the metabolism of drugs by the different CYP450 and UGT isoforms. Important factors for these differences are differences in activity, e.g. due to genetic polymorphisms, and the amount of protein of the respective enzyme in the liver.
The aim of this study was to provide a systematic overview of the variability in protein levels of CYP450 and UGT isoforms in the human liver. To this end, a manually curated database of protein abundance information was established and used to answer the following questions: (i) What are the protein distributions of the different CYP450 and UGT isoforms? (ii) What factors affect these protein levels (age, sex, body mass index, smoking, alcohol consumption)? (iii) How do CYP and UGT isoforms correlate among each other?
Key results include: (i) A free and open database of CYP and UGT protein levels in the human liver has been established. (ii) CYP and UGT isoforms show large inter-individual variability in protein levels. (iii) Different CYP and UGT isoforms have different protein levels. (iv) Factors such as age, smoking and alcohol consumption may affect the protein levels of CYP and UGT enzymes. (v) There are significant correlations in protein levels between CYP isoforms and between UGT isoforms.
This work has established an important resource for the study of variability in drug detoxification due to variability in protein levels in the human liver.
Keywords:
cytochrome P450,
UDP-glucuronosyltransferase,
CYP,
UGT,
protein,
liver,
|
|
| 2024 |


|
Bayesian modelling of time series data (BayModTS)-a FAIR workflow to process sparse and highly variable data
Höpfl S, Albadry M, Dahmen U, Herrmann KH, Kindler EM, König M, Reichenbach JR, Tautenhahn HM, Wei W, Zhao WT, Radde NE.
Bioinformatics. 2024 May 2;40(5):btae312 (publication). doi:10.1093/bioinformatics/btae312. pmid:38741151
Motivation: Systems biology aims to better understand living systems through mathematical modelling of experimental and clinical data. A pervasive challenge in quantitative dynamical modelling is the integration of time series measurements, which often have high variability and low sampling resolution. Approaches are required to utilize such information while consistently handling uncertainties. Results: We present BayModTS (Bayesian modelling of time series data), a new FAIR (findable, accessible, interoperable, and reusable) workflow for processing and analysing sparse and highly variable time series data. BayModTS consistently transfers uncertainties from data to model predictions, including process knowledge via parameterized models. Further, credible differences in the dynamics of different conditions can be identified by filtering noise. To demonstrate the power and versatility of BayModTS, we applied it to three hepatic datasets gathered from three different species and with different measurement techniques: (i) blood perfusion measurements by magnetic resonance imaging in rat livers after portal vein ligation, (ii) pharmacokinetic time series of different drugs in normal and steatotic mice, and (iii) CT-based volumetric assessment of human liver remnants after clinical liver resection. Availability and implementation: The BayModTS codebase is available on GitHub at https://github.com/Systems-Theory-in-Systems-Biology/BayModTS. The repository contains a Python script for the executable BayModTS workflow and a widely applicable SBML (systems biology markup language) model for retarded transient functions. In addition, all examples from the paper are included in the repository. Data and code of the application examples are stored on DaRUS: https://doi.org/10.18419/darus-3876. The raw MRI ROI voxel data were uploaded to DaRUS: https://doi.org/10.18419/darus-3878. The steatosis metabolite data are published on FairdomHub: 10.15490/fairdomhub.1.study.1070.1.
Keywords:
Animals,
Bayes Theorem,
Humans,
Liver metabolism,
Magnetic Resonance Imaging,
Mice,
Rats,
Software,
Systems Biology,
|
|
| 2024 |



|
The simulation experiment description markup language (SED-ML): language specification for level 1 version 5
Smith LP, Bergmann FT, Garny A, Helikar T, Karr J, Nickerson D, Sauro H, Waltemath D, König M.
J Integr Bioinform. 2024 Apr 15 (publication). doi:10.1515/jib-2024-0008. pmid:38613325
Modern biological research is increasingly informed by computational simulation experiments, which necessitate the development of methods for annotating, archiving, sharing, and reproducing the conducted experiments. These simulations increasingly require extensive collaboration among modelers, experimentalists, and engineers. The Minimum Information About a Simulation Experiment (MIASE) guidelines outline the information needed to share simulation experiments. SED-ML is a computer-readable format for the information outlined by MIASE, created as a community project and supported by many investigators and software tools. Level 1 Version 5 of SED-ML expands the ability of modelers to define simulations in SED-ML using the Kinetic Simulation Algorithm Onotoloy (KiSAO). While it was possible in Version 4 to define a simulation entirely using KiSAO, Version 5 now allows users to define tasks, model changes, ranges, and outputs using the ontology as well. SED-ML is supported by a growing ecosystem of investigators, model languages, and software tools, including various languages for constraint-based, kinetic, qualitative, rule-based, and spatial models, and many simulation tools, visual editors, model repositories, and validators. Additional information about SED-ML is available at https://sed-ml.org/.
Keywords:
SED-ML,
Standardization,
|
|
| 2024 |


|
Simulation of zonation-function relationships in the liver using coupled multiscale models: Application to drug-induced liver injury
Steffen Gerhäusser, Lena Lambers, Luis Mandl, Julian Franquinet, Tim Ricken, Matthias König
bioRxiv 2024.03.26.586870; (preprint). doi:10.1101/2024.03.26.586870
Multiscale modeling requires the coupling of models on different scales, often based on different mathematical approaches and developed by different research teams. This poses many challenges, such as defining interfaces for coupling, reproducible exchange of submodels, efficient simulation of the models, or reproducibility of results. Here, we present a multiscale digital twin of the liver that couples a partial differential equation (PDE)-based porous media approach for the hepatic lobule with cellular-scale ordinary differential equation (ODE)-based models. The models based on the theory of porous media describe transport, tissue mechanical properties, and deformations at the lobular scale, while the cellular models describe hepatic metabolism in terms of drug metabolism and damage in terms of necrosis. The resulting multiscale model of the liver was used to simulate perfusion-zonation-function relationships in the liver spanning scales from single cell to the lobulus. The model was applied to study the effects of (i) protein zonation patterns (metabolic zonation) and (ii) drug concentration dependence on spatially heterogeneous liver damage in the form of necrosis. Depending on the zonation pattern, different liver damage patterns could be reproduced, including periportal and pericentral necrosis as seen in drug-induced liver injury (DILI). Increasing the drug concentration led to an increase in the observed damage pattern. A key point for the success was the integration of domain-specific simulators based on standard exchange formats, i.e., libroadrunner for the high-performance simulation of ODE-based systems and FEBio for the simulation of the continuum-biomechanical part. This allows a standardized and reproducible exchange of cellular scale models in the Systems Biology Markup Language (SBML) between research groups.
Keywords:
liver,
computational model,
biomechanics,
porous media,
drug metabolism,
digital twin,
DILI,
cytochrome P450,
|
|
| 2024 |

|
Quantifying fat zonation in liver lobules: an integrated multiscale in silico model combining disturbed microperfusion and fat metabolism via a continuum biomechanical bi-scale, tri-phasic approach
Lena Lambers, Navina Waschinsky, Jana Schleicher, Matthias König, Hans-Michael Tautenhahn, Mohamed Albadry, Uta Dahmen and Tim Ricken
Biomech Model Mechanobiol. 2024 Feb 25. (publication). doi:10.1007/s10237-023-01797-0. pmid:38402347
Metabolic zonation refers to the spatial separation of metabolic functions along the sinusoidal axes of the liver. This phenomenon forms the foundation for adjusting hepatic metabolism to physiological requirements in health and disease (e.g., metabolic dysfunction-associated steatotic liver disease/MASLD). Zonated metabolic functions are influenced by zonal morphological abnormalities in the liver, such as periportal fibrosis and pericentral steatosis. We aim to analyze the interplay between microperfusion, oxygen gradient, fat metabolism and resulting zonated fat accumulation in a liver lobule. Therefore we developed a continuum biomechanical, tri-phasic, bi-scale, and multicomponent in silico model, which allows to numerically simulate coupled perfusion-function-growth interactions two-dimensionally in liver lobules. The developed homogenized model has the following specifications: (i) thermodynamically consistent, (ii) tri-phase model (tissue, fat, blood), (iii) penta-substances (glycogen, glucose, lactate, FFA, and oxygen), and (iv) bi-scale approach (lobule, cell). Our presented in silico model accounts for the mutual coupling between spatial and time-dependent liver perfusion, metabolic pathways and fat accumulation. The model thus allows the prediction of fat development in the liver lobule, depending on perfusion, oxygen and plasma concentration of free fatty acids (FFA), oxidative processes, the synthesis and the secretion of triglycerides (TGs). The use of a bi-scale approach allows in addition to focus on scale bridging processes. Thus, we will investigate how changes at the cellular scale affect perfusion at the lobular scale and vice versa. This allows to predict the zonation of fat distribution (periportal or pericentral) depending on initial conditions, as well as external and internal boundary value conditions.
Keywords:
Multiscale Approach,
Homogenization,
Theory of Porous Media (TPM),
Growth Kinematics,
Metabolic dysfunction-Associated Steatotic Liver Disease (MASLD),
FEM boundary problem,
|
|
| 2024 |

|
SimLivA – Modelling ischemia reperfusion injury in the liver: A first step towards a clinical decision support tool.
Hans-Michael Tautenhahn, Tim Ricken, Uta Dahmen, Luis Mandl, Laura Bütow, Steffen Gerhäusser, Lena Lambers, Xinpei Chen, Elina Lehmann, Olaf Dirsch, Matthias König
GAMM-Mitteilungen. (2024), e202370003 (proceeding). doi:10.1002/gamm.202370003
The SIMulation supported LIVer Assessment for donor organs (SimLivA) project aims to develop a mathematical model to accurately simulate the influence of mechanical alterations in marginal liver grafts (specifically steatotic ones) and cold ischemia on early ischemia-reperfusion injury (IRI) during liver transplantation. Our project tackles significant research challenges, including the co-development of computational methodologies, experimental studies, clinical processes, and technical workflows. We aim to refine a continuum-biomechanical model for enhanced IRI prediction, collect pivotal experimental and clinical data, and assess the clinical applicability of our model. Our efforts involve augmenting and tailoring a coupled continuum-biomechanical, multiphase, and multi-scale partial differential equation-ordinary differential equation (PDE-ODE) model of the liver lobule, allowing us to numerically simulate IRI depending on the degree of steatosis and the duration of ischemia. The envisaged model will intertwine the structure, perfusion, and function of the liver, serving as a crucial aid in clinical decision-making processes. We view this as the initial step towards an in-silico clinical decision support tool aimed at enhancing the outcomes of liver transplantation. In this paper, we provide an overview of the SimLivA project and our preliminary findings, which include: a cellular model that delineates critical processes in the context of IRI during transplantation; and the integration of this model into a multi-scale PDE-ODE model using a homogenized, multi-scale, multi-component approach within the Theory of Porous Media (TPM) framework. The model has successfully simulated the interconnected relationship between structure, perfusion, and function—all of which are integral to IRI. Initial results show simulations at the cellular scale that describe critical processes related to IRI during transplantation. After integrating this model into a multiscale PDE-ODE model, first simulations were performed on the spatial distribution of key functions during warm and cold ischaemia. In addition, we were able to study the effect of tissue perfusion and temperature, two critical parameters in the context of liver transplantation and IRI.
Keywords:
ischemia-reperfusion injury,
IRI,
liver transplantation,
ODE,
PDE,
porous media,
|
|
| 2024 |


|
A physiologically-based pharmacokinetic (PBPK) model of sorafenib for investigating the effect of sorafenib parameters in hepatorenal impairment
Internship Report Frances Okibedi (supervisor: Matthias König)
Internship report, July 2024 (report)
Liver cancer ranks second in cancer-related deaths thereby making timely diagnosis and personalized treatment crucial. Hepatocellular carcinoma (HCC) patients often get diagnosed at advanced stages, hindering curative options like resection and ablation. Systemic treatment has become the standard therapy and sorafenib has been the primary systemic treatment for advanced HCC for the last ten years, being the first approved drug for this purpose. Sorafenib (Nexavar), is a multi-kinase inhibitor medication approved for primarily treating advanced renal cell carcinoma, hepatocellular carcinoma or other cancers. After oral administration, sorafenib enters hepatocytes via OATP1B-type carriers. It undergoes CYP3A4-mediated oxidative metabolism to sorafenib-N-oxide (M2) and UGT1A9-mediated glucuronidation forming sorafenib-glucuronide (SG). SG is secreted into bile mainly by ABCC2, with a fraction reuptaken by ABCC3, preventing ABCC2 saturation. In bile, SG can be excreted via the feces or converted back to sorafenib by a bacterial β-glucuronidase. Sorafenib is absorbed in the intestine, reentering the systemic circulation, contributing to elimination and detoxification. Sorafenib is excreted mainly by the feces (77%) and urine (19%). Hepatic and renal impairment could have a large impact on the pharmacokinetics of sorafenib. Within this work the pharmacokinetics of sorafenib were analyzed by developing a physiologically-based pharmacokinetic (PBPK) model based on extensive data curation of sorafenib data. The model allowed to simulate the time-concentration courses of sorafenib in various tissues and to calculate the pharmacokinetic parameters for sorafenib under single and multiple dosing regimes. The model was applied to study the dose-dependency of sorafenib treatment and to investigate the effect of hepatic and renal impairment on sorafenib pharmacokinetics.
Keywords:
sorafenib,
PBPK,
pharmacokinetics,
HCC,
liver function,
renal function,
|
|
| 2023 |

|
Eleven Strategies for Making Reproducible Research and Open Science Training the Norm at Research Institutions
Friederike E Kohrs, Susann Auer, Alexandra Bannach-Brown, Susann Fiedler, Tamarinde Laura Haven, Verena Heise, Constance Holman, Flavio Azevedo, René Bernard, Armin Bleier, Nicole Bössel, Brian Patrick Cahill, Leyla Jael Castro, Adrian Ehrenhofer, Kristina Eichel, Maximillian Frank, Claudia Frick, Malte Friese, Anne Gärtner, Kerstin Gierend, David Joachim Grüning, Lena Hahn, Maren Hülsemann, Malika Ihle, Sabrina Illius, Laura König, Matthias König, Louisa Kulke, Anton Kutlin, Fritjof Lammers, David MA Mehler, Christoph Miehl, Anett Müller-Alcazar, Claudia Neuendorf, Helen Niemeyer, Florian Pargent, Aaron Peikert, Christina U Pfeuffer, Robert Reinecke, Jan Philipp Röer, Jessica L Rohmann, Alfredo Sánchez-Tójar, Stefan Scherbaum, Elena Sixtus, Lisa Spitzer, Vera Maren Straßburger, Marcel Weber, Clarissa J Whitmire, Josephine Zerna, Dilara Zorbek, Philipp Zumstein, Tracey L Weissgerber
eLife (2023) 12:e89736. (publication). doi:10.7554/eLife.89736
Reproducible research and open science practices have the potential to accelerate scientific progress by allowing others to reuse research outputs, and by promoting rigorous research that is more likely to yield trustworthy results. However, these practices are uncommon in many fields, so there is a clear need for training that helps and encourages researchers to integrate reproducible research and open science practices into their daily work. Here, we outline eleven strategies for making training in these practices the norm at research institutions. The strategies, which emerged from a virtual brainstorming event organized in collaboration with the German Reproducibility Network, are concentrated in three areas: (i) adapting research assessment criteria and program requirements; (ii) training; (iii) building communities. We provide a brief overview of each strategy, offer tips for implementation, and provide links to resources. We also highlight the importance of allocating resources and monitoring impact. Our goal is to encourage researchers – in their roles as scientists, supervisors, mentors, instructors, and members of curriculum, hiring or evaluation committees – to think creatively about the many ways they can promote reproducible research and open science practices in their institutions.
Keywords:
reproducible research,
open science,
training,
|
|
| 2023 |


|
Insights into Intestinal P-glycoprotein Function using Talinolol: A PBPK Modeling Approach
Beatrice Stemmer Mallol, Jan Grzegorzewski, Matthias König
bioRxiv 2023.11.21.568168 (preprint). doi:10.1101/2023.11.21.568168
Talinolol is a cardioselective beta-blocker used in the treatment of various cardiovascular diseases and tachyarrhythmias. Its intestinal absorption is determined by uptake by the organic anion transporting polypeptide 2B1 (OATP2B1) and efflux via P-glycoprotein (P-gp). Talinolol can be taken up via OATP1B1 in the liver, where it enters the enterohepatic circulation. Talinolol is excreted unchanged in the urine and feces. Talinolol is widely used as a probe drug for the intestinal efflux transporter P-gp, which plays a critical role in protecting against potentially toxic substances and facilitating the elimination of xenobiotics. In this work, an extensive database of talinolol pharmacokinetics was established and used to develop and validate a physiologically based pharmacokinetic (PBPK) model of talinolol for P-gp phenotyping. The model was used to investigate the influence of several factors on talinolol pharmacokinetics: (i) inhibition of P-gp via drug-drug interaction; (i) genetic polymorphisms of P-gp; (iii) activity of OATP2B1 and OATP1B1; (iv) effect of disease, namely hepatic and renal impairment; and (v) site-specific distribution of P-gp and OATP2B1 in the intestine. The model accurately predicts the concentration-time profile of talinolol after oral and intravenous administration of single and multiple dosing. Furthermore, the model accurately describes the effect of genetic variants of P-gp on the pharmacokinetics of talinolol, the effect of inhibition of P-gp, the effect of renal impairment, as well as site-specific infusion of talinolol in the intestine. The detailed description of the intestinal absorption of talinolol and the predictions of talinolol pharmacokinetics as a function of hepatorenal impairment provide valuable clinical insights for metabolic phenotyping with talinolol. Both the model and the database are freely available for reuse.
Keywords:
PBPK,
talinolol,
P-glycoprotein,
pharmacokinetics,
OATP2B1,
OATB1B1,
renal impairment,
cirrhosis,
|
|
| 2023 |


|
A pathway model of glucose-stimulated insulin secretion in the pancreatic β-cell
Maheshvare MD., Raha S., König M.Δ, and Pal D.Δ (Δ equal contribution)
Front. Endocrinol. 14:1185656 (publication). doi:10.3389/fendo.2023.1185656
The pancreas plays a critical role in maintaining glucose homeostasis through the secretion of hormones from the islets of Langerhans. Glucose-stimulated insulin secretion (GSIS) by the pancreatic β-cell is the main mechanism for reducing elevated plasma glucose. Here we present a systematic modeling workflow for the development of kinetic pathway models using the Systems Biology Markup Language (SBML). Steps include retrieval of information from databases, curation of experimental and clinical data for model calibration and validation, integration of heterogeneous data including absolute and relative measurements, unit normalization, data normalization, and model annotation. An important factor was the reproducibility and exchangeability of the model, which allowed the use of various existing tools. The workflow was applied to construct a novel data-driven kinetic model of GSIS in the pancreatic β-cell based on experimental and clinical data from 39 studies spanning 50 years of pancreatic, islet, and β-cell research in humans, rats, mice, and cell lines. The model consists of detailed glycolysis and phenomenological equations for insulin secretion coupled to cellular energy state, ATP dynamics and (ATP/ADP ratio). Key findings of our work are that in GSIS there is a glucose-dependent increase in almost all intermediates of glycolysis. This increase in glycolytic metabolites is accompanied by an increase in energy metabolites, especially ATP and NADH. One of the few decreasing metabolites is ADP, which, in combination with the increase in ATP, results in a large increase in ATP/ADP ratios in the β-cell with increasing glucose. Insulin secretion is dependent on ATP/ADP, resulting in glucose-stimulated insulin secretion. The observed glucose-dependent increase in glycolytic intermediates and the resulting change in ATP/ADP ratios and insulin secretion is a robust phenomenon observed across data sets, experimental systems and species. Model predictions of the glucose-dependent response of glycolytic intermediates and biphasic insulin secretion are in good agreement with experimental measurements. Our model predicts that factors affecting ATP consumption, ATP formation, hexokinase, phosphofructokinase, and ATP/ADP-dependent insulin secretion have a major effect on GSIS. In conclusion, we have developed and applied a systematic modeling workflow for pathway models that allowed us to gain insight into key mechanisms in GSIS in the pancreatic β-cell.
Keywords:
glucose-stimulated insulin secretion,
GSIS,
glycolysis,
pancreas,
kinetic model,
systems biology,
|
|
| 2023 |


|
A physiologically based pharmacokinetic (PBPK) model of the probe drug talinolol for the characterization of intestinal P-glycoprotein
Bachelor Thesis Beatrice Stemmer Mallol (supervisor: Matthias König)
Bachelor Thesis, July 2023 (thesis)
Talinolol is a cardioselective beta-blocker used for the treatment of various cardiovascular diseases and tachyarrhythmia. The gastrointestinal absorption of talinolol is determined via its uptake in the intestine via the organic anion transporting polypeptide 2B1 (OATP2B1) and its efflux via the P-glycoprotein (P-gp). After intestinal absorption talinolol can be transported into the liver via OATP1B1 talinolol where it undergoes enterohepatic circulation. Talinolol is excreted unchanged in the urine and feces. In addition to its clinical application, talinolol is widely used as a probe drug for the intestinal efflux transporter P-glycoprotein. P-gp plays a crucial role in the human body as it is expressed in various tissues to protect against potentially toxic substances, facilitating the elimination of xenobiotics. The application of talinolol for P-gp phenotyping enables evaluation of factors influencing P-gp-mediated transport in vivo such as genetic polymorphisms of P-gp as well as the distribution of P-gp along the intestine. Within this thesis, an extensive dataset of talinolol pharmacokinetics was established and used to develop a physiologically based pharmacokinetic (PBPK) model for talinolol. The model was applied to investigate the influence of various factors on the pharmacokinetics of talinolol, including: (i) genetic variants of P-gp; (ii) enzymatic activity of the transporters OATP2B1 and OATP1B1; (iii) site-specific distribution of P-gp and OATP2B1 proteins in the intestine, and (iv) the impact of diseases such as liver cirrhosis and renal dysfunction. The model enables precise predictions of the concentration-time profile of talinolol in various tissues following oral or intravenous administration. Furthermore, the model accurately describes the effect of genetic variants of P-gp on the pharmacokinetics of talinolol. The detailed description of the limiting intestinal absorption of intestinal provided by the model, along with the precise prediction of talinolol`s pharmacokinetics in different renal functions, holds significant clinical relevance.
Keywords:
talinolol,
PBPK,
pharmacokinetics,
OATP2B1,
OATP1B1,
P-glycoprotein,
liver function,
renal function,
|
|
| 2023 |


|
A physiologically based pharmacokinetic model for CYP2E1 phenotyping via chlorzoxazone
J. Küttner, J. Grzegorzewski, HM. Tautenhahn, M. König
bioRxiv 2023.04.12.536571 (preprint). doi:10.1101/2023.04.12.536571
The cytochrome P450 (CYP) superfamily of enzymes plays a critical role in the metabolism of drugs, toxins, and endogenous and exogenous compounds. The activity of CYP enzymes can be influenced by a variety of factors, including genetics, diet, age, environmental factors, and disease. Among the major isoforms, CYP2E1 is of particular interest due to its involvement in the metabolism of various low molecular weight chemicals, including alcohols, pharmaceuticals, industrial solvents, and halogenated anesthetics. Metabolic phenotyping of CYPs based on the elimination of test compounds is a useful method for assessing in vivo activity, with chlorzoxazone being the primary probe drug for phenotyping of CYP2E1. The aim of this work was to investigate the effect of changes in CYP2E1 level and activity, ethanol consumption, ethanol abstinence, and liver impairment on the results of metabolic phenotyping with chlorzoxazone. To accomplish this, an extensive pharmacokinetic dataset for chlorzoxazone was established and a physiologically based pharmacokinetic (PBPK) model of chlorzoxazone and its metabolites, 6-hydroxychlorzoxazone and chlorzoxazone-O-glucuronide, was developed and validated. The model incorporates the effect of ethanol consumption on CYP2E1 levels and activity by extending the model with a core ethanol pharmacokinetic model and a CYP2E1 turnover model. The model accurately predicts pharmacokinetic data from several clinical studies and is able to estimate the effect of changes in CYP2E1 levels and activity on chlorzoxazone pharmacokinetics. Regular ethanol consumption induces CYP2E1 over two to three weeks, resulting in increased conversion of chlorzoxazone to 6-hydroxychlorzoxazone and a higher 6-hydroxychlorzoxazone/chlorzoxazone metabolic ratio. After ethanol withdrawal, CYP2E1 levels return to baseline within one week. Importantly, liver impairment has an opposite effect, resulting in reduced liver function via CYP2E1. In alcoholics with liver impairment who also consume ethanol, these factors will have opposite confounding effects on metabolic phenotyping with chlorzoxazone.
Keywords:
PBPK,
chlorzoxazone,
CYP2E1,
alcohol,
ethanol,
pharmacokinetics,
|
|
| 2023 |


|
Specifications of Standards in Systems and Synthetic Biology: Status and Developments in 2022 and the COMBINE meeting 2022
M. König, P. Gleeson, M. Golebiewski, T. Gorochowski, M. Hucka, S. Keating, C. Myers, D. Nickerson, F. Schreiber
J Integr Bioinform. 2023 Mar 29;20(1). (publication). doi:10.1515/jib-2023-0004. pmid:36989443
This special issue of the Journal of Integrative Bioinformatics contains updated specifications of COMBINE standards in systems and synthetic biology. The 2022 special issue presents three updates to the standards: CellML 2.0.1, SBML Level 3 Package: Spatial Processes, Version 1, Release 1, and Synthetic Biology Open Language (SBOL) Version 3.1.0. This document can also be used to identify the latest specifications for all COMBINE standards. In addition, this editorial provides a brief overview of the COMBINE 2022 meeting in Berlin.
Keywords:
COMBINE,
standardization,
SBML,
SEDML,
OMEX,
|
|
| 2023 |


|
standard-GEM: standardization of open-source genome-scale metabolic models
Mihail Anton, Eivind Almaas, Rui Benfeitas, Sara Benito-Vaquerizo, Lars M. Blank, Andreas Dräger, John M. Hancock, Cheewin Kittikunapong, Matthias König, Feiran Li, Ulf W. Liebal, Hongzhong Lu, Hongwu Ma, Radhakrishnan Mahadevan, Adil Mardinoglu, Jens Nielsen, Juan Nogales, Marco Pagni, Jason A. Papin, Kiran Raosaheb Patil, Nathan D. Price, Jonathan L. Robinson, Benjamín J. Sánchez, Maria Suarez-Diez, Snorre Sulheim, L. Thomas Svensson, Bas Teusink, Wanwipa Vongsangnak, Hao Wang, Ahmad A. Zeidan, Eduard J. Kerkhoven
bioRxiv 2023.03.21.512712 (preprint). doi:10.1101/2023.03.21.512712
The field of metabolic modelling at the genome-scale continues to grow with more models being created and curated. This comes with an increasing demand for adopting common principles regarding transparency and versioning, in addition to standardisation efforts regarding file formats, annotation and testing. Here, we present a standardised template for git-based and GitHub-hosted genome-scale metabolic models (GEMs) supporting both new models and curated ones, following FAIR principles (findability, accessibility, interoperability, and reusability), and incorporating best-practices. standard-GEM facilitates the reuse of GEMs across web services and platforms in the metabolic modelling field and enables automatic validation of GEMs. The use of this template for new models, and its adoption for existing ones, paves the way for increasing model quality, openness, and accessibility with minimal effort.
Keywords:
genome-scale metabolic model,
open source,
versioning,
|
|
| 2023 |


|
Simvastatin therapy in different subtypes of hypercholesterolemia - a physiologically based modelling approach
F. Bartsch, J. Grzegorzewski, H. Pujol, HM. Tautenhahn, M. König
medRxiv 2023.02.01.23285358 (preprint). doi:10.1101/2023.02.01.23285358
Hypercholesterolemia is a multifaceted plasma lipid disorder with heterogeneous causes including lifestyle and genetic factors. A key feature of hypercholesterolemia is elevated plasma levels of low-density lipoprotein cholesterol (LDL-C). Several genetic variants have been reported to be associated with hypercholesterolemia, known as familial hypercholesterolemia (FH). Important variants affect the LDL receptor (LDLR), which mediates the uptake of LDL-C from the plasma, apoliporotein B (APOB), which is involved in the binding of LDL-C to the LDLR, and proprotein convertase subtilisin/kexin type 9 (PCSK9), which modulates the degradation of the LDLR. A typical treatment for hypercholesterolemia is statin medication, with simvastatin being one of the most commonly prescribed statins. In this work, the LDL-C lowering therapy with simvastatin in hypercholesterolemia was investigated using a computational modeling approach. A physiologically based pharmacokinetic model of simvastatin integrated with a pharmacodynamic model of plasma LDL-C (PBPK/PD) was developed based on extensive data curation. A key component of the model is LDL-C turnover by the liver, consisting of: hepatic cholesterol synthesis with the key enzymes HMG-CoA reductase and HMG-CoA synthase; cholesterol export from the liver as VLDL-C; de novo synthesis of LDLR; transport of LDLR to the membrane; binding of LDL-C by LDLR via APOB; endocytosis of the LDLR-LDL-C complex; recycling of LDLR from the complex. The model was applied to study the effects of simvastatin therapy in hypercholesterolemia due to different causes in the LDLR pathway corresponding to different subtypes of hypercholesterolemia. Model predictions of LDL-C lowering therapy were validated with independent clinical data sets. Key findings are: (i) hepatic LDLR turnover is highly heterogeneous among FH classes; (ii) despite this heterogeneity, simvastatin therapy results in a consistent reduction in plasma LDL-C regardless of class; and (iii) simvastatin therapy shows a dose-dependent reduction in LDL-C. Our model suggests that the underlying cause of hypercholesterolemia does not influence simvastatin therapy. Furthermore, our model supports the treatment strategy of stepwise dose adjustment to achieve target LDL-C levels. Both the model and the database are freely available for reuse.
Keywords:
simvastatin,
hypercholesterolemia,
LDL-cholesterol,
LDL-receptor,
physiologically based pharmacokinetic model,
pharmacokinetics,
pharmacodynamics,
PK/PD,
|
|
| 2023 |


|
Physiologically based pharmacokinetic (PBPK) modeling for dynamical liver function tests and CYP phenotyping
Jan Grzegorzewski (supervisor: Matthias König)
PhD Thesis, Jan 2023 (thesis)
Cytochrome P450 (CYP) phenotyping and dynamic liver function testing are essential methods in clinical practice. These methods utilize the pharmacokinetics (PK) of test substances and their metabolites to gain insight into the liver’s metabolic capacity and the activity of enzymes and transporters. Despite an extensive body of literature, many aspects affecting liver function and CYP activity are not well understood. Liver function tests are not only influenced by numerous characteristics of a studied subject but also by the specifics of individual study procedures. A key challenge is to disentangle the various factors which influence the outcome of the measurements from each other to study their influence on the dynamic liver function and CYP phenotype. In this work, the challenge was addressed through meta-analysis and physiologically based pharmacokinetic (PBPK) modeling. As a foundation, an open pharmacokinetics database (https://pk-db.com) was developed and pharmacokinetics data were curated for a wide range of test substances. To my knowledge, PK-DB currently contains the largest open pharmacokinetic dataset on substances used for phenotyping and dynamical liver function testing. The dataset allowed for identifying and quantifying demographic and racial bias (sex, ethnicity, age, health), reporting errors, and inconsistencies in pharmacokinetic literature. Based on the data, a caffeine pharmacokinetics meta-analysis was conducted concerning various factors affecting liver function and CYP1A2 activity. In particular, meta-analysis and data integration solidified existing knowledge on the effects of smoking, oral contraceptives, multiple diseases, and co-medications on caffeine pharmacokinetics. Similarly, the measurement accuracy of caffeine concentration in saliva versus plasma was quantified, and the effect of dosing amount and sampling timing for phenotyping were analyzed. In addition, the impact of CYP2D6 polymorphism was investigated. Therefore, a PBPK model of dextromethorphan (DXM) and its metabolites dextrorphan (DXO) and dextrorphan O-glucuronide (DXO-Glu) was developed, and calibrated and validated with pharmacokinetics data. The variability in CYP activity was modeled based on in vitro data. The model can predict individual plasma concentrations and urinary amounts of DXM, DXO, and DXO-Glu and the metabolic phenotype based on the individual’s CYP2D6 genotype and physiological characteristics. The analyses suggest that most of the variability in the pharmacokinetics of dextromethorphan can be attributed to the variability in CYP2D6 and CYP3A4 enzyme kinetics. Among various other investigations, the influence of ethnicity on CYP2D6 activity was also investigated. Contributions to PK data curation and PBPK model development were also made for other phenotyping and liver function test substances (chlorzoxazone, codeine, diazepam, galactose, indocyanine green (ICG), metoprolol, midazolam, omeprazole, pravastatin, simvastatin, talinolol, and torasemide). For ICG, in particular, the impact of hepatic blood flow, cardiac output, and body weight, as well as the survival probability after partial hepatectomy, were investigated by PBPK modeling. Notably, the studying of the various test substances was only made possible by a systematic and standardized workflow that facilitated data integration, data sharing, the creation of reproducible PBPK models, and the standardized integration of data and models. In conclusion, a pharmacokinetic database, methods, and workflows for the analysis of test compounds used in dynamical liver function testing and CYP phenotyping were established. Factors affecting CYP phenotyping and liver function testing were investigated by meta-analysis and PBPK modeling. The models developed in this effort have the potential to impact personalized medicine and to increase the precision of dynamic liver function tests in clinics.
Keywords:
liver function,
metabolic phenotyping,
pharmacokinetics,
PBPK,
|
|
| 2023 |



|
libRoadRunner 2.0: A High-Performance SBML Simulation and Analysis Library
Ciaran Welsh, Jin Xu, Lucian Smith, Matthias König, Kiri Choi, Herbert M. Sauro
Bioinformatics. 2023 Jan 1;39(1):btac770 (publication). doi:10.1093/bioinformatics/btac770. pmid:36478036
Motivation: This paper presents libRoadRunner 2.0, an extensible, high-performance, cross-platform, open-source software library for the simulation and analysis of models expressed using the Systems Biology Markup Language (SBML). Results: libRoadRunner is a self-contained library, able to run either as a component inside other tools via its C ++, C and Python APIs, or interactively through its Python or Julia interface. libRoadRunner uses a custom Just-In-Time (JIT) compiler built on the widely-used LLVM JIT compiler framework. It compiles SBML-specified models directly into native machine code for a large variety of processors, making it fast enough to simulate extremely large models or repeated runs in reasonable timeframes. libRoadRunner is flexible, supporting the bulk of the SBML specification (except for delay and nonlinear algebraic equations) as well as several SBML extensions such as hierarchical composition and probability distributions. It offers multiple deterministic and stochastic integrators, as well as tools for steady-state, sensitivity, stability and structural analyses. Availability: libRoadRunner binary distributions for Windows, Mac OS, and Linux, Julia and Python bindings, source code, and documentation are all available at https://github.com/sys-bio/roadrunner, and Python bindings are also available via pip. The source code can be compiled for the supported systems as well as in principle any system supported by LLVM-13, such as ARM-based computers like the Raspberry Pi. The library is licensed under the Apache License Version 2.0.
Keywords:
computer simulation,
software,
systems biology,
|
|
| 2022 |

|
Cytochrome p450 enzymes in periportal steatosis: Modulation of drug metabolizing activity but not of pericentral expression pattern
M. Albadry, S. Höpfl, N. Ehteshamzad, M. König, M. Böttcher, J. Neumann, A. Lupp, O. Dirsch, N. Radde, B. Christ, M. Christ, L.O. Schwen, H. Laue, R. Klopfleisch and Uta Dahmen
Sci Rep. 2022 Dec 17;12(1):21825 (publication). doi:10.1038/s41598-022-26483-6. pmid:36528753
Little is known about the impact of morphological disorders in distinct zones on metabolic zonation. It was described recently that periportal fibrosis did affect the expression of CYP proteins, a set of pericentrally located drug-metabolizing enzymes. Here, we investigated whether periportal steatosis might have a similar effect. Periportal steatosis was induced in C57BL6/J mice by feeding a high-fat diet with low methionine/choline content for either two or four weeks. Steatosis severity was quantified using image analysis. Triglycerides and CYP activity were quantified in photometric or fluorometric assay. The dis- tribution of CYP3A4, CYP1A2, CYP2D6, and CYP2E1 was visualized by immunohistochemistry. Pharmacokinetic parameters of test drugs were determined after injecting a drug cocktail (caffeine, codeine, and midazolam). The dietary model resulted in moderate to severe mixed steatosis confined to periportal and midzonal areas. Periportal steatosis did not affect the zonal distribution of CYP expression but the activity of selected CYPs was associated with steatosis severity. Caffeine elimination was accelerated by microvesicular steatosis, whereas midazolam elimination was delayed in macrovesicular steatosis. In summary, periportal steatosis affected parameters of pericentrally located drug metabolism. This observation calls for fur- ther investigations of the highly complex interrelationship between steatosis and drug metabolism and underlying signaling mechanisms.
Keywords:
cytochrome P-450 Enzyme System,
fatty Liver,
mice,
pharmacokinetics,
|
|
| 2022 |


|
Physiologically based pharmacokinetic (PBPK) modeling of the role of CYP2D6 polymorphism for metabolic phenotyping with dextromethorphan
Grzegorzewski, J., Brandhorst, J., König, M.
Front Pharmacol. 2022 Oct 24;13:1029073 (publication). doi:10.3389/fphar.2022.1029073. pmid:36353484
The cytochrome P450 2D6 (CYP2D6) is a key xenobiotic-metabolizing enzyme involved in the clearance of many drugs. Genetic polymorphisms in CYP2D6 contribute to the large inter-individual variability in drug metabolism and could affect metabolic phenotyping of CYP2D6 probe substances such as dextromethorphan (DXM). To study this question, we (i) established an extensive pharmacokinetics dataset for DXM; and (ii) developed and validated a physiologically based pharmacokinetic (PBPK) model of dextromethorphan (DXM) and its metabolites dextrorphan (DXO) and dextrorphan O-glucuronide (DXO-Glu) based on the data. Drug-gene interactions (DGI) were introduced by accounting for changes in CYP2D6 enzyme kinetics depending on activity score (AS), which in combination with AS for individual polymorphisms allowed us to model of CYP2D6 gene variants. Variability in CYP3A4 and CYP2D6 activity was modeled based on in vitro data from human liver microsomes. Model predictions are in very good agreement with pharmacokinetics data for CYP2D6 polymorphisms, CYP2D6 activity as measured by AS, and CYP2D6 metabolic phenotypes (UM, EM, IM, PM). The model was applied to investigate the genotype-phenotype association and the role of CYP2D6 polymorphisms for metabolic phenotyping with dextromethorphan using the urinary cumulative metabolic ratio DXM/(DXO+DXO-Glu) (UCMR). The effect of parameters on UCMR was studied via sensitivity analysis. Model predictions indicate very good robustness against the intervention protocol (i.e. application form, dosing amount, dissolution rate, and sampling time) and good robustness against physiological variation. The model is capable of estimating the UCMR dispersion within and across populations depending on activity scores. Moreover, the distribution of UCMR and the risk of genotype-phenotype mismatch could be estimated for populations with known CYP2D6 genotype frequencies. The model can be applied as a tool for individual prediction of UCMR and metabolic phenotype based on CYP2D6 genotype. Both, model and database are freely available for reuse.
Keywords:
Dextromethorphan,
CYP2D6,
Physiologically based pharmacokinetic model,
PBPK,
Pharmacokinetics,
Pharmacogenomics,
|
|
| 2022 |



|
FAIR Sharing of Reproducible Models of Epidemic and Pandemic Forecast
Ramachandran, K.Δ; König, M.Δ; Scharm, M.; Nguyen, T.V.N.; Hermjakob, H.; Waltemath, D.; Malik Sheriff, R.S. (Δ equal contribution)
Preprints 2022, 2022060137 (preprint). doi:10.20944/preprints202206.0137.v1
A major challenge for the dissemination, replication, and reuse of epidemiological forecasting studies during COVID-19 pandemics is the lack of clear guidelines and platforms to exchange models in a Findable, Accessible, Interoperable, and Reusable (FAIR) manner, facilitating reproducibility of research outcomes. During the beginning of pandemics, models were developed in diverse tools that were not interoperable, opaque without traceability and semantics, and scattered across various platforms - making them hard to locate, infer and reuse. In this work, we demonstrate that implementing the standards developed by the systems biology community to encode and share COVID-19 epidemiological models can serve as a roadmap to implement models as a tool in medical informatics, in general. As a proof-of-concept, we encoded and shared 24 epidemiological models using the standard format for model exchange in systems biology, annotated them with cross-references to data resources, packed up all associated files in COMBINE archives for easy sharing, and finally, disseminated the models through BioModels repository to significantly enhance their reproducibility and repurposing potential. We recommend the use of systems biology standards to encode and share models of epidemic and pandemic forecasts to improve their findability, accessibility, interoperability, reusability, and reproducibility.
Keywords:
FAIR,
epidemiology,
models,
pandemic forecast,
SIR modelling,
standards,
|
|
| 2022 |



|
BioSimulators: a central registry of simulation engines and services for recommending specific tools
Shaikh B, Smith LP, Vasilescu D, Marupilla G, Wilson M, Agmon E, Agnew H, Andrews SS, Anwar A, Beber ME, Bergmann FT, Brooks D, Brusch L, Calzone L, Choi K, Cooper J, Detloff J, Drawert B, Dumontier M, Ermentrout GB, Faeder JR, Freiburger AP, Fröhlich F, Funahashi A, Garny A, Gennari JH, Gleeson P, Goelzer A, Haiman Z, Hasenauer J, Hellerstein JL, Hermjakob H, Hoops S, Ison JC, Jahn D, Jakubowski HV, Jordan R, Kalaš M, König M, Liebermeister W, Sheriff RSM, Mandal S, McDougal R, Medley JK, Mendes P, Müller R, Myers CJ, Naldi A, Nguyen TVN, Nickerson DP, Olivier BG, Patoliya D, Paulevé L, Petzold LR, Priya A, Rampadarath AK, Rohwer JM, Saglam AS, Singh D, Sinha A, Snoep J, Sorby H, Spangler R, Starruß J, Thomas PJ, van Niekerk D, Weindl D, Zhang F, Zhukova A, Goldberg AP, Schaff JC, Blinov ML, Sauro HM, Moraru II, Karr JR.
Nucleic Acids Res. 2022 May 7;50(W1):W108–14 (publication). doi:10.1093/nar/gkac331. pmid:35524558
Computational models have great potential to accelerate bioscience, bioengineering, and medicine. However, it remains challenging to reproduce and reuse simulations, in part, because the numerous formats and methods for simulating various subsystems and scales remain siloed by different software tools. For example, each tool must be executed through a distinct interface. To help investigators find and use simulation tools, we developed BioSimulators (https://biosimulators.org), a central registry of the capabilities of simulation tools and consistent Python, command-line and containerized interfaces to each version of each tool. The foundation of BioSimulators is standards, such as CellML, SBML, SED-ML and the COMBINE archive format, and validation tools for simulation projects and simulation tools that ensure these standards are used consistently. To help modelers find tools for particular projects, we have also used the registry to develop recommendation services. We anticipate that BioSimulators will help modelers exchange, reproduce, and combine simulations.
Keywords:
reproducibility,
software,
simulation,
|
|
| 2022 |


|
A physiologically based model of pravastatin - The role of genotypes and hepatic or renal impairment on the pharmacokinetics of pravastatin
Helena Leal Pujol (supervisor: Matthias König)
Bachelor Thesis, May 2022 (thesis)
Hypercholesterolaemia, i.e., elevated plasma levels of cholesterol, is a major risk factor for cardiovascular disease, the leading cause of death globally. Hypercholesterolaemia can be treated using statins, a class of medications which inhibit HMG-CoA reductase, a major enzyme in cholesterol synthesis. Pravastatin is a statin used to reduce total and low-density plasma cholesterol levels and increase high-density plasma cholesterol levels in hypercholesterolaemic patients. Pravastatin is absorbed from the small intestine by the transporter OATP2B1 and subsequently transported in the liver via OATP1B1 from where it can be exported in the bile via the enzymatic exporter MRP2. Pravastatin can be excreted either in the urine via the kidneys or in the faeces due to incomplete absorption. Hepatic and renal impairment could have a large impact on the pharmacokinetics of statins as could have genetic variants of the transporters OATP2B1, OATP1B1 and MRP2. Within this thesis the pharmacokinetics of pravastatin were analysed by developing a physiologically-based pharmacokinetics model based on extensive data curation of pravastatin data. The model allows to simulate the time-concentration courses of pravastatin in various tissues and to calculate the pharmacokinetic parameters for pravastatin. Furthermore, the model was applied to investigate the effects of genotypes of the enzymatic transporters and hepatic and renal impairment on the pharmacokinetics of pravastatin. Thus, key questions such as, how pravastatin therapy would be affected in renal or hepatic disease, as well as how pravastatin therapy should be adapted based on genotypes, find an answer in this work.
Keywords:
pravastatin,
PBPK,
pharmacokinetics,
OATP2B1,
OATP1B1,
MRP2,
liver function,
renal function,
|
|
| 2022 |



|
Dynamic Flux Balance Analysis Models in SBML
Matthias KönigΔ, Leandro H. WatanabeΔ, Jan Grzegorzewski, and Chris J. Myers. (Δ equal contribution)
bioRxiv 245076 (preprint). doi:10.1101/245076
Computational models in systems biology and systems medicine are typically simulated using a single formalism such as ordinary differential equations (ODE). However, more complex models require the coupling of multiple formalisms since different biological phenomena are better described by different methods. For example, metabolism in steady state is often modeled using flux-balance analysis (FBA) whereas dynamic changes of model components are better described via ODEs. The coupling of FBA and ODE modeling formalisms results in dynamic FBA models. A major challenge is how to describe such hybrid models that couple multiple formalisms in a standardized way so that they can be exchanged between tools and simulated consistently in a reproducible manner. This paper presents a scheme for encoding and implementation of dynamic FBA models in the Systems Biology Markup Language (SBML), thereby enabling the exchange of multi-framework computational models between software tools. We demonstrate the feasibility of the approach using various example models and show that different tools are able to simulate the hybrid models and agree on the results. As part of this work, two independent implementations of a multi-framework simulation method for dynamic FBA have been developed supporting such models: iBioSim and sbmlutils.
Keywords:
dynamic flux balance analysis,
DFBA,
flux balance analysis (FBA),
ordinary differential equations (ODE),
static optimization approach (SOA),
|
|
| 2022 |


|
Pharmacokinetics of caffeine: A systematic analysis of reported data for application in metabolic phenotyping and liver function testing
Jan Grzegorzewski, Florian Bartsch, Adrian Köller, and Matthias König
Frontiers in Pharmacology 2022, Vol12 (publication). doi:10.3389/fphar.2021.752826. pmid:35280254
Caffeine is by far the most ubiquitous psychostimulant worldwide found in tea, coffee, cocoa, energy drinks, and many other beverages and food. Caffeine is almost exclusively metabolized in the liver by the cytochrome P-450 enzyme system to the main product paraxanthine and the additional products theobromine and theophylline. Besides its stimulating properties, two important applications of caffeine are metabolic phenotyping of cytochrome P450 1A2 (CYP1A2) and liver function testing. An open challenge in this context is to identify underlying causes of the large inter-individual variability in caffeine pharmacokinetics. Data is urgently needed to understand and quantify confounding factors such as lifestyle (e.g., smoking), the effects of drug-caffeine interactions (e.g., medication metabolized via CYP1A2), and the effect of disease. Here we report the first integrative and systematic analysis of data on caffeine pharmacokinetics from 141 publications and provide a comprehensive high-quality data set on the pharmacokinetics of caffeine, caffeine metabolites, and their metabolic ratios in human adults. The data set is enriched by meta-data on the characteristics of studied patient cohorts and subjects (e.g., age, body weight, smoking status, health status), the applied interventions (e.g., dosing, substance, route of application), measured pharmacokinetic time-courses, and pharmacokinetic parameters (e.g., clearance, half-life, area under the curve). We demonstrate via multiple applications how the data set can be used to solidify existing knowledge and gain new insights relevant for metabolic phenotyping and liver function testing based on caffeine. Specifically, we analyzed 1) the alteration of caffeine pharmacokinetics with smoking and use of oral contraceptives; 2) drug-drug interactions with caffeine as possible confounding factors of caffeine pharmacokinetics or source of adverse effects; 3) alteration of caffeine pharmacokinetics in disease; and 4) the applicability of caffeine as a salivary test substance by comparison of plasma and saliva data. In conclusion, our data set and analyses provide important resources which could enable more accurate caffeine-based metabolic phenotyping and liver function testing.
Keywords:
CYP1A2,
caffeine,
drug-disease interactions,
drug-drug interactions,
liver function test,
oral contraceptives,
pharmacokinetics,
smoking,
|
|
| 2021 |


|
Prediction of survival after hepatectomy using a physiologically based pharmacokinetic model of indocyanine green liver function tests
Adrian Köller, Jan Grzegorzewski, Michael Tautenhahn, Matthias König
Front. Physiol., 22 November 2021 (publication). doi:10.3389/fphys.2021.730418. pmid:34880771
The evaluation of hepatic function and functional capacity of the liver are essential tasks in hepatology as well as in hepatobiliary surgery. Indocyanine green (ICG) is a widely applied test compound that is used in clinical routine to evaluate hepatic function. Important questions for the functional evaluation with ICG in the context of hepatectomy are how liver disease such as cirrhosis alters ICG elimination, and if postoperative survival can be predicted from preoperative ICG measurements. Within this work a physiologically based pharmacokinetic (PBPK) model of ICG was developed and applied to the prediction of the effects of a liver resection under various degrees of cirrhosis. For the parametrization of the computational model and validation of model predictions a database of ICG pharmacokinetic data was established. The model was applied (i) to study the effect of liver cirrhosis and liver resection on ICG pharmacokinetics; and (ii) to evaluate the model-based prediction of postoperative ICG-R15 (retention ratio 15 min after administration) as a measure for postoperative outcome. Key results are the accurate prediction of changes in ICG pharmacokinetics caused by liver cirrhosis and postoperative changes of ICG-elimination after liver resection, as validated with a wide range of data sets. Based on the PBPK model, individual survival after liver resection could be classified, demonstrating its potential value as a clinical tool.
Keywords:
computational model,
hepatectomy,
indocyanine green,
liver cirrhosis,
liver function,
liver resection,
mathematical model,
pharmacokinetics,
|
|
| 2021 |


|
Physiologically based modeling of the effect of physiological and anthropometric variability on indocyanine green based liver function tests
Adrian Köller, Jan Grzegorzewski and Matthias König
Front Physiol. 2021 Nov 22;12:757293 (publication). doi:10.3389/fphys.2021.757293. pmid:34880776
Accurate evaluation of liver function is a central task in hepatology. Dynamic liver function tests (DLFT) based on the time-dependent elimination of a test substance provide an important tool for such a functional assessment. These tests are used in the diagnosis and monitoring of liver disease as well as in the planning of hepatobiliary surgery. A key challenge in the evaluation of liver function with DLFTs is the large inter-individual variability. Indocyanine green (ICG) is a widely applied test compound used for the evaluation of liver function. After an intravenous administration, pharmacokinetic (PK) parameters are calculated from the plasma disappearance curve of ICG which provide an estimate of liver function. The hepatic elimination of ICG is affected by physiological factors such as hepatic blood flow or binding of ICG to plasma proteins, anthropometric factors such as body weight, age, and sex, or the protein amount of the organic anion-transporting polypeptide 1B3 (OATP1B3) mediating the hepatic uptake of ICG. Being able to account for and better understand these various sources of inter-individual variability would allow to improve the power of ICG based DLFTs and move toward an individualized evaluation of liver function. Within this work we systematically analyzed the effect of various factors on ICG elimination by the means of computational modeling. For the analysis, a recently developed and validated physiologically based pharmacokinetics (PBPK) model of ICG distribution and hepatic elimination was utilized. Key results are (i) a systematic analysis of the variability in ICG elimination due to hepatic blood flow, cardiac output, OATP1B3 abundance, liver volume, body weight and plasma bilirubin level; (ii) the evaluation of the inter-individual variability in ICG elimination via a large in silico cohort of n = 100,000 subjects based on the NHANES cohort with special focus on stratification by age, sex, and body weight; (iii) the evaluation of the effect of various degrees of cirrhosis on variability in ICG elimination. The presented results are an important step toward individualizing liver function tests by elucidating the effects of confounding physiological and anthropometric parameters in the evaluation of liver function via ICG.
Keywords:
ICG,
PBPK,
computational model,
indocyanine green,
liver cirrhosis,
liver function,
mathematical model,
mathematical model,
pharmacokinetics,
|
|
| 2021 |


|
Hepatectomy-Induced Alterations in Hepatic Perfusion and Function - Toward Multi-Scale Computational Modeling for a Better Prediction of Post-hepatectomy Liver Function
Bruno ChristΔ, Maximilian CollatzΔ, Uta DahmenΔ, Karl-Heinz HerrmannΔ, Sebastian HöpflΔ, Matthias KönigΔ, Lena LambersΔ, Manja MarzΔ, Daria MeyerΔ, Nicole RaddeΔ, Jürgen R. ReichenbachΔ, Tim RickenΔ, and Hans-Michael TautenhahnΔ (Δ equal contribution)
Front Physiol. 2021 Nov 18;12:733868 (review). doi:10.3389/fphys.2021.733868. pmid:34867441
Liver resection causes marked perfusion alterations in the liver remnant both on the organ scale (vascular anatomy) and on the microscale (sinusoidal blood flow on tissue level). These changes in perfusion affect hepatic functions via direct alterations in blood supply and drainage, followed by indirect changes of biomechanical tissue properties and cellular function. Changes in blood flow impose compression, tension and shear forces on the liver tissue. These forces are perceived by mechanosensors on parenchymal and non-parenchymal cells of the liver and regulate cell-cell and cell-matrix interactions as well as cellular signaling and metabolism. These interactions are key players in tissue growth and remodeling, a prerequisite to restore tissue function after PHx. Their dysregulation is associated with metabolic impairment of the liver eventually leading to liver failure, a serious post-hepatectomy complication with high morbidity and mortality. Though certain links are known, the overall functional change after liver surgery is not understood due to complex feedback loops, non-linearities, spatial heterogeneities and different time-scales of events. Computational modeling is a unique approach to gain a better understanding of complex biomedical systems. This approach allows (i) integration of heterogeneous data and knowledge on multiple scales into a consistent view of how perfusion is related to hepatic function; (ii) testing and generating hypotheses based on predictive models, which must be validated experimentally and clinically. In the long term, computational modeling will (iii) support surgical planning by predicting surgery-induced perfusion perturbations and their functional (metabolic) consequences; and thereby (iv) allow minimizing surgical risks for the individual patient. Here, we review the alterations of hepatic perfusion, biomechanical properties and function associated with hepatectomy. Specifically, we provide an overview over the clinical problem, preoperative diagnostics, functional imaging approaches, experimental approaches in animal models, mechanoperception in the liver and impact on cellular metabolism, omics approaches with a focus on transcriptomics, data integration and uncertainty analysis, and computational modeling on multiple scales. Finally, we provide a perspective on how multi-scale computational models, which couple perfusion changes to hepatic function, could become part of clinical workflows to predict and optimize patient outcome after complex liver surgery.
Keywords:
liver perfusion,
liver regeneration,
liver surgery,
mechanoperception,
multi-scale modeling,
perfusion/function relationship,
rodent models of liver surgery,
|
|
| 2021 |


|
Specifications of Standards in Systems and Synthetic Biology: Status and Developments in 2021
F. Schreiber, P. Gleeson, M. Golebiewski, T. Gorochowski, M. Hucka, S. Keating, M. König, C. Myers, D. Nickerson, D. Walthemath
J Integr Bioinform. 2021 Oct 22;18(3):20210026 (publication). doi:10.1515/jib-2021-0026. pmid:34674411
This special issue of the Journal of Integrative Bioinformatics contains updated specifications of COMBINE standards in systems and synthetic biology. The 2021 special issue presents four updates of standards: Synthetic Biology Open Language Visual Version 2.3, Synthetic Biology Open Language Visual Version 3.0, Simulation Experiment Description Markup Language Level 1 Version 4, and OMEX Metadata specification Version 1.2. This document can also be consulted to identify the latest specifications of all COMBINE standards.
Keywords:
COMBINE,
standardization,
SBML,
SEDML,
OMEX,
|
|
| 2021 |


|
Computational modelling of omeprazole - pharmacokinetics and pharmacodynamics
Sükrü Balci (supervisor: Matthias König)
Bachelor Thesis, October 2021 (thesis)
Proton pump inhibitors (PPIs) are one of the most commonly used drugs whose main action is to inhibit the formation of gastric acid. Among the PPIs, omeprazole is one of the most important and most prescribed. Omeprazole is rapidly absorbed by the stomach and has a short half-life of around 1-2 hours in plasma. Omeprazole reduces gastric acid secretion via non-competitive selective inhibition of the H+/K+-ATPases in the gastric parietal cells, resulting in an increase in stomach pH. After cessation of omeprazole treatment, it takes a few days until the gastric acid secretion returns to baseline due to protein turnover of the proton pumps. Repeated dosing with omeprazole results in a dose-potentiation due to an increased absorption of omeprazole with increasing pH. Within this thesis, the pharmacokinetics and pharmacodynamics of omeprazole were studied using a computational modeling approach. Based on extensive curation of clinical data of omeprazole a physiological-based pharmacokinetics/pharmacodynamics model of omeprazole and its effect on stomach pH was developed. The model allowed to simulate the pharmacokinetics of omeprazole under varying doses and treatment regimens and to study the resulting inhibition of the proton pumps and changes in stomach pH. Key questions studied in this thesis were (i) how gastric acid output and pH change with omeprazole dosing and treatment regime, and ii) what the effects of repeated dosing on omeprazole absorption and pH are.
Keywords:
omeprazol,
PBPK,
pharmacokinetics,
pharmacodynamics,
proton pump inhibitor,
PPI,
|
|
| 2021 |



|
SBMLWebApp: Web-based Simulation, Steady-State Analysis, and Parameter Estimation of Systems Biology Models
Takahiro G. Yamada, Kaito Ii, Matthias König, Martina Feierabend, Andreas Dräger, and Akira Funahashi
Processes 9, no. 10: 1830 (publication). doi:10.3390/pr9101830
In systems biology, biological phenomena are often modeled by Ordinary Differential Equations (ODEs) and distributed in the de facto standard file format SBML. The primary analyses performed with such models are dynamic simulation, steady-state analysis, and parameter estimation. These methodologies are mathematically formalized, and libraries for such analyses have been published. Several tools exist to create, simulate, or visualize models encoded in SBML. However, setting up and establishing analysis environments is a crucial hurdle for non-modelers. Therefore, easy access to perform fundamental analyses of ODE models is a significant challenge. We developed SBMLWebApp, a web-based service to execute SBML-based simulation, steady-state analysis, and parameter estimation directly in the browser without the need for any setup or prior knowledge to address this issue. SBMLWebApp visualizes the result and numerical table of each analysis and provides a download of the results. SBMLWebApp allows users to select and analyze SBML models directly from the BioModels Database. Taken together, SBMLWebApp provides barrier-free access to an SBML analysis environment for simulation, steady-state analysis, and parameter estimation for SBML models. SBMLWebApp is implemented in Java™ based on an Apache Tomcat® web server using COPASI, the Systems Biology Simulation Core Library (SBSCL), and LibSBMLSim as simulation engines. SBMLWebApp is licensed under MIT with source code freely available. At the end of this article, the Data Availability Statement gives the internet links to the two websites to find the source code and run the program online.
Keywords:
SBML,
kinetic models,
time-course simulation,
steady-state simulation,
parameter estimation,
model calibration,
software,
web application,
|
|
| 2021 |



|
The simulation experiment description markup language (SED-ML): language specification for level 1 version 4
Smith, Lucian P., Bergmann, Frank T., Garny, Alan, Helikar, Tomáš, Karr, Jonathan, Nickerson, David, Sauro, Herbert, Waltemath, Dagmar and König, Matthias.
Journal of Integrative Bioinformatics, vol. 18, no. 3, 2021, pp. 20210021 (publication). doi:10.1515/jib-2021-0021. pmid:35330701
Computational simulation experiments increasingly inform modern biological research, and bring with them the need to provide ways to annotate, archive, share and reproduce the experiments performed. These simulations increasingly require extensive collaboration among modelers, experimentalists, and engineers. The Minimum Information About a Simulation Experiment (MIASE) guidelines outline the information needed to share simulation experiments. SED-ML is a computer-readable format for the information outlined by MIASE, created as a community project and supported by many investigators and software tools. The first versions of SED-ML focused on deterministic and stochastic simulations of models. Level 1 Version 4 of SED-ML substantially expands these capabilities to cover additional types of models, model languages, parameter estimations, simulations and analyses of models, and analyses and visualizations of simulation results. To facilitate consistent practices across the community, Level 1 Version 4 also more clearly describes the use of SED-ML constructs, and includes numerous concrete validation rules. SED-ML is supported by a growing ecosystem of investigators, model languages, and software tools, including eight languages for constraint-based, kinetic, qualitative, rule-based, and spatial models, over 20 simulation tools, visual editors, model repositories, and validators. Additional information about SED-ML is available at https://sed-ml.org/.
Keywords:
COMBINE,
SED-ML,
|
|
| 2021 |



|
The Systems Biology Simulation Core Library
Panchiwala, H.; Shah, S.; Planatscher, H.; Zakharchuk, M.; König, M.; Dräger, A.
Bioinformatics. 2022 Jan 12;38(3):864-865 (publication). doi:10.1093/bioinformatics/btab669. pmid:34554191
Summary: Studying biological systems generally relies on computational modeling and simulation, e.g., model-driven discovery and hypothesis testing. Progress in standardization efforts led to the development of interrelated file formats to exchange and reuse models in systems biology, such as SBML, the Simulation Experiment Description Markup Language (SED-ML) or the Open Modeling EXchange format. Conducting simulation experiments based on these formats requires efficient and reusable implementations to make them accessible to the broader scientific community and to ensure the reproducibility of the results. The Systems Biology Simulation Core Library (SBSCL) provides interpreters and solvers for these standards as a versatile open-source API in JavaTM. The library simulates even complex bio-models and supports deterministic Ordinary Differential Equations; Stochastic Differential Equations; constraint-based analyses; recent SBML and SED-ML versions; exchange of results, and visualization of in silico experiments; open modeling exchange formats (COMBINE archives); hierarchically structured models; and compatibility with standard testing systems, including the Systems Biology Test Suite and published models from the BioModels and BiGG databases.
|
|
| 2021 |

|
Ten Simple Rules for FAIR Sharing of Experimental and Clinical Data with the Modeling Community
Matthias KönigΔ, Jan Grzegorzewski, Martin Golebiewski, Henning Hermjakob, Mike Hucka, Brett Olivier, Sarah M. Keating, David Nickerson, Falk Schreiber, Rahuman Sheriff, Dagmar Waltemath
Preprints 2021, 2021080303 (preprint). doi:10.20944/preprints202108.0303.v2
Science continues to become more interdisciplinary and to involve increasingly complex data sets. Many projects in the biomedical and health-related sciences follow or aim to follow the principles of FAIR data sharing, which has been demonstrated to foster collaboration, to lead to better research outcomes, and to help ensure reproducibility of results. Data generated in the course of biomedical and health research present specific challenges for FAIR sharing in the sense that they are heterogeneous and highly sensitive to context and the needs of protection and privacy. Data sharing must respect these features without impeding timely dissemination of results, so that they can contribute to time-critical advances in medical therapy and treatment. Modeling and simulation of biomedical processes have become established tools, and a global community has been developing algorithms, methodologies, and standards for applying biomedical simulation models in clinical research. However, it can be difficult for clinician scientists to follow the specific rules and recommendations for FAIR data sharing within this domain. We seek to clarify the standard workflow for sharing experimental and clinical data with the simulation modeling community. By following these recommendations, data sharing will be improved, collaborations will become more effective, and the FAIR publication and subsequent reuse of data will become possible at the level of quality necessary to support biomedical and health-related sciences.
Keywords:
data sharing,
FAIR,
|
|
| 2021 |

|
OMEX Metadata Specification Version 1.2
John Gennari, Matthias König, Goskel Misirli, Maxwell Neal, David Nickerson, Dagmar Waltemath
J Integr Bioinform. 2021 Oct 20;18(3):20210020 (publication). doi:10.1515/jib-2021-0020. pmid:34668356
A standardized approach to annotating computational biomedical models and their associated files can facilitate model reuse and reproducibility among research groups, enhance search and retrieval of models and data, and enable semantic comparisons between models. Motivated by these potential benefits and guided by consensus across the COmputational Modeling in BIology NEtwork (COMBINE) community, we have developed a specification for encoding annotations in Open Modeling and EXchange (OMEX)-formatted archives. This document details version 1.2 of the specification, which builds on version 1.0 published last year in this journal. In particular, this version includes a set of initial model-level annotations (whereas v 1.0 described exclusively annotations at a smaller scale). Additionally, this version uses best practices for namespaces, and introduces omex-library.org as a common root for all annotations. Distributing modeling projects within an OMEX archive is a best practice established by COMBINE, and the OMEX metadata specification presented here provides a harmonized, community-driven approach for annotating a variety of standardized model representations. This specification acts as a technical guideline for developing software tools that can support this standard, and thereby encourages broad advances in model reuse, discovery, and semantic analyses.
|
|
| 2021 |



|
SED-ML Validator: tool for debugging simulation experiments
Bilal Shaikh, Andrew Philip Freiburger, Matthias König, Frank T. Bergmann, David P. Nickerson, Herbert M. Sauro, Michael L. Blinov, Lucian P. Smith, Ion I. Moraru, Jonathan R. Karr
arXiV, 2021 (preprint). doi:10.48550/arXiv.2106.00844
|
|
| 2021 |

|
A Physiologically Based Model of Indocyanine Green Liver Function Tests - Effects of Physiological Factors, Hepatic Disease and Hepatic Surgery
Adrian Köller (supervisor: Matthias König)
Bachelor Thesis, March 2021 (thesis)
The evaluation of hepatic function and functional capacity are essential tasks in hepatology. Indocyanine Green (ICG) is a widely applied test compound that is used in clinical routine to evaluate hepatic function. Accurate assessment of liver-function through ICG pharmacokinetics is challenging because the elimination of ICG is influenced by physiological factors such as liver perfusion as well as the health status of the patient. Important questions for the functional evaluation are how blood flow, plasma protein binding or surgical interventions can influence the distribution and elimination of ICG. In the course of this project, these questions were studied by the means of computational modeling to evaluate ICG dynamics in silico. Within this work a physiological-based model of ICG pharmacokinetics was developed. For the parameterization and validation of the model a database of ICG pharmacokinetic data was established. The model was applied to study (i) the effect of liver disease, specifically liver cirrhosis, on ICG pharmacokinetics; (ii) how hepatic blood flow and other cardiovascular parameters affect ICG-elimination; and (iii) the role of plasma protein binding of ICG and its impact on ICG pharmacokinetics. Finally, the model was applied to analyse the effect of liver resection (hepatectomy) on ICG-elimination and the value of ICG as a predictive measure for postoperative outcome. The model is able to accurately predict changes in ICG pharmacokinetics caused by changes in blood flow and plasma proteins as well as liver cirrhosis. Furthermore, the model is able to accurately predict postoperative changes of ICG-elimination after liver resection, showing its potential value as a clinical tool.
|
|
| 2020 |


|
QuaLiPerF – Multi-X Liver Modelling
Tim Ricken, Lena Lambers, Bruno Christ, Uta Dahmen, Karl-Heinz Herrmann, Matthias König, Manja Marz, Nicole Radde, Jürgen R. Reichenbach, Lars Ole Schwen & Hans-Michael Tautenhahn
GACM - German Association for Computational Mechanics Report, winter 2020: No. 13 (proceeding)
|
|
| 2020 |

|
Computational Modelling of Simvastatin - Effects on HMG-CoA Reductase Activity and Cholesterol
Florian Bartsch (supervisor: Matthias König)
Bachelor Thesis, November 2020 (thesis)
Cholesterol is one of the most important molecules in biology. Elevated plasma levels are associated with an increased risk for arteriosclerosis and cardiovascular diseases. A common treatment for hypercholesterolemic patients are statins, which are capable of reducing total plasma cholesterol levels. Within this thesis the effects of the statin simvastatin on cholesterol levels were studied using a computational modelling approach. Based on extensive data curation, a kinetic model of simvastatin pharmacokinetics coupled to a pharmacodynamic model of cholesterol metabolism was developed. The resulting pharmacokinetic/ pharmacodynamic (PK/PD) model was applied to study the effects of simvastatin on hepatic HMG-CoA reductase activity, hepatic cholesterol homeostasis and plasma cholesterol levels.
Keywords:
simvastaton,
PBPK,
pharmacokinetics,
pharmacodynamics,
cholesterol,
|
|
| 2020 |

|
Computational Modelling of Midazolam Clearance: Effect of Inhibitors and Inducers
Yannick Duport (supervisor: Matthias König)
Bachelor Thesis, August 2020 (thesis)
Midazolam is one of the most effective and safe medicines used for anesthesia and procedural sedation, as well as treating trouble sleeping and severe agitation. Midazolam is metabolised in the liver as well as in the small intestine to cytochrome P450 3A4 (CYP3A4). Many factors can alter the pharmacokinetics of midazolam due to inhibition or induction of hepatic and/or intestinal metabolism, resulting in either (i) adverse effects due to increased plasma midazolam concentrations in case of inhibition of metabolism or (ii) ineffectiveness as a consequence of too low concentrations if metabolism is induced. Within this thesis the clearance and pharmacokinetics of midazolam were studied, using a computational modeling approach. A large data base of midazolam pharmacokinetics data was established and utilized to develop a physiological-based pharmacokinetics model of midazolam. The model was applied to study the following questions: (i) What are the effects of CYP3A4 inhibition and induction on midazolam pharmacokinetics? (ii) What are the differences of intestinal and hepatic CYP3A4 inhibition and induction on midazolam pharmacokinetics?
Keywords:
midazolam,
PBPK,
pharmacokinetics,
drug-drug interactions,
DDI,
|
|
| 2020 |



|
PK-DB: pharmacokinetics database for individualized and stratified computational modeling
Grzegorzewski J, Brandhorst J, Green K, Eleftheriadou D, Duport Y, Barthorscht F, Köller A, Ke DYJ, De Angelis S, König M.
Nucleic Acids Res. 2021 Jan 8;49(D1):D1358-D1364 (publication). doi:10.1093/nar/gkaa990. pmid:33151297
A multitude of pharmacokinetics studies have been published. However, due to the lack of an open database, pharmacokinetics data, as well as the corresponding meta-information, have been difficult to access. We present PK-DB (https://pk-db.com), an open database for pharmacokinetics information from clinical trials. PK-DB provides curated information on (i) characteristics of studied patient cohorts and subjects (e.g. age, bodyweight, smoking status, genetic variants); (ii) applied interventions (e.g. dosing, substance, route of application); (iii) pharmacokinetic parameters (e.g. clearance, half-life, area under the curve) and (iv) measured pharmacokinetic time-courses. Key features are the representation of experimental errors, the normalization of measurement units, annotation of information to biological ontologies, calculation of pharmacokinetic parameters from concentration-time profiles, a workflow for collaborative data curation, strong validation rules on the data, computational access via a REST API as well as human access via a web interface. PK-DB enables meta-analysis based on data from multiple studies and data integration with computational models. A special focus lies on meta-data relevant for individualized and stratified computational modeling with methods like physiologically based pharmacokinetic (PBPK), pharmacokinetic/pharmacodynamic (PK/PD), or population pharmacokinetic (pop PK) modeling.
|
|
| 2020 |



|
SBML Level 3: an extensible format for the exchange and reuse of biological models
SM Keating, D Waltemath, M König, F Zhang, A Dräger, C Chaouiya, FT Bergmann, A Finney, CS Gillespie, T Helikar, S Hoops, RS Malik-Sheriff, SL Moodie, II Moraru, CJ Myers, A Naldi, BG Olivier, S Sahle, JC Schaff, LP Smith, MJ Swat, DT, L Watanabe, DJ Wilkinson, ML Blinov, K Begley, JR Faeder, HF Gómez, TM Hamm, Y Inagaki, W Liebermeister, AL Lister, D Lucio, E Mjolsness, CJ Proctor, K Raman, N Rodriguez, CA Shaffer, BE Shapiro, J Stelling, N Swainston, N Tanimura, J Wagner, M Meier-Schellersheim, HM Sauro, B Palsson, H Bolouri, H Kitano, Akira Funahashi, H Hermjakob, JC Doyle M Hucka, and SBML Community members
Mol Syst Biol. 2020;16(8):e9110 (publication). doi:10.15252/msb.20199110. pmid:32845085
Systems biology has experienced dramatic growth in the number, size, and complexity of computational models. To reproduce simulation results and reuse models, researchers must exchange unambiguous model descriptions. We review the latest edition of the Systems Biology Markup Language (SBML), a format designed for this purpose. A community of modelers and software authors developed SBML Level 3 over the past decade. Its modular form consists of a core suited to representing reaction-based models and packages that extend the core with features suited to other model types including constraint-based models, reaction-diffusion models, logical network models, and rule-based models. The format leverages two decades of SBML and a rich software ecosystem that transformed how systems biologists build and interact with models. More recently, the rise of multiscale models of whole cells and organs, and new data sources such as single-cell measurements and live imaging, has precipitated new ways of integrating data with models. We provide our perspectives on the challenges presented by these developments and how SBML Level 3 provides the foundation needed to support this evolution.
Keywords:
computational modeling,
file format,
interoperability,
reproducibility,
systems biology,
|
|
| 2020 |


|
Open modeling and exchange (OMEX) metadata specification version 1.0
Neal ML, Gennari JH, Waltemath D, Nickerson DP, König M
J Integr Bioinform. 2020 Jun 25;17(2-3):20200020 (publication). doi:10.1515/jib-2020-0020. pmid:32589606
A standardized approach to annotating computational biomedical models and their associated files can facilitate model reuse and reproducibility among research groups, enhance search and retrieval of models and data, and enable semantic comparisons between models. Motivated by these potential benefits and guided by consensus across the COmputational Modeling in BIology NEtwork (COMBINE) community, we have developed a specification for encoding annotations in Open Modeling and EXchange (OMEX)-formatted archives. Distributing modeling projects within these archives is a best practice established by COMBINE, and the OMEX metadata specification presented here provides a harmonized, community-driven approach for annotating a variety of standardized model and data representation formats within an archive. The specification primarily includes technical guidelines for encoding archive metadata, so that software tools can more easily utilize and exchange it, thereby spurring broad advancements in model reuse, discovery, and semantic analyses.
Keywords:
data integration,
metadata,
semantics,
simulation,
|
|
| 2020 |


|
Specifications of Standards in Systems and Synthetic Biology:Status and Developments in 2020
F. Schreiber, B. Sommer, G. Bader, P. Gleeson, M. Golebiewski, T. Gorochowski, M. Hucka, S. Keating, M. König, C. Myers, D. Nickerson, D. Walthemath
J Integr Bioinform. 2020 Jun 29;17(2-3):20200022 (publication). doi:10.1515/jib-2020-0022
This special issue of the Journal of Integrative Bioinformatics presents papers related to the 10th COMBINE meeting together with the annual update of COMBINE standards in systems and synthetic biology.
Keywords:
ontologies,
standards,
synthetic biology,
systems biology,
|
|
| 2020 |


|
The first 10 years of the international coordination network for standards in systems and synthetic biology (COMBINE)
D Waltemath, M Golebiewski, M Blinov, P Gleeson, H Hermjakob, E Inau, S Keating, M König, O Krebs, R Malik-Sheriff, D Nickerson, E Oberortner, H Sauro, F Schreiber, L Smith, M Stefan, U Wittig, C Myers
J Integr Bioinform. 2020 Jun 29;17(2-3):20200005 (publication). doi:10.1515/jib-2020-0005
This paper presents a report on outcomes of the 10th Computational Modeling in Biology Network (COMBINE) meeting that was held in Heidelberg, Germany, in July of 2019. The annual event brings together researchers, biocurators and software engineers to present recent results and discuss future work in the area of standards for systems and synthetic biology. The COMBINE initiative coordinates the development of various community standards and formats for computational models in the life sciences. Over the past 10 years, COMBINE has brought together standard communities that have further developed and harmonized their standards for better interoperability of models and data. COMBINE 2019 was co-located with a stakeholder workshop of the European EU-STANDS4PM initiative that aims at harmonized data and model standardization for in silico models in the field of personalized medicine, as well as with the FAIRDOM PALs meeting to discuss findable, accessible, interoperable and reusable (FAIR) data sharing. This report briefly describes the work discussed in invited and contributed talks as well as during breakout sessions. It also highlights recent advancements in data, model, and annotation standardization efforts. Finally, this report concludes with some challenges and opportunities that this community will face during the next 10 years.
Keywords:
COMBINE,
community building,
meeting report,
standardization,
|
|
| 2020 |


|
The Distributions Package for SBML Level 3
L Smith, S Moodie, F Bergmann, C Gillespie, S Keating, M. König, C Myers, M Swat, D Wilkinson, M Hucka
J Integr Bioinform. 2020 Jul 20;17(2-3):20200018 (publication). doi:10.1515/jib-2020-0018. pmid:32750035
Biological models often contain elements that have inexact numerical values, since they are based on values that are stochastic in nature or data that contains uncertainty. The Systems Biology Markup Language (SBML) Level 3 Core specification does not include an explicit mechanism to include inexact or stochastic values in a model, but it does provide a mechanism for SBML packages to extend the Core specification and add additional syntactic constructs. The SBML Distributions package for SBML Level 3 adds the necessary features to allow models to encode information about the distribution and uncertainty of values underlying a quantity.
Keywords:
SBML,
distributions,
modeling,
systems biology,
uncertainty,
|
|
| 2020 |



|
MEMOTE for standardized genome-scale metabolic model testing
Christian Lieven, Moritz Emanuel Beber, Brett G. Olivier, Frank T. Bergmann, Meric Ataman, Parizad Babaei, Jennifer A. Bartell, Lars M. Blank, Siddharth Chauhan, Kevin Correia, Christian Diener, Andreas Dräger, Birgitta Elisabeth Ebert, Janaka N. Edirisinghe, Jose P. Faria, Adam Feist, Georgios Fengos, Ronan M. T. Fleming, Beatriz Garcia-Jimenez, Vassily Hatzimanikatis, Wout van Helvoirt, Christopher Henry, Henning Hermjakob, Markus J. Herrgard, Hyun Uk Kim, Zachary King, Jasper Jan Koehorst, Matthias König, Steffen Klamt, Edda Klipp, Meiyappan Lakshmanan, Nicolas Le Novere, Dong-Yup Lee, Sang Yup Lee, Sunjae Lee, Nathan E. Lewis, Hongwu Ma, Daniel Machado, Radhakrishnan Mahadevan, Paulo Maia, Adil Mardinoglu, Greg L. Medlock, Jonathan Monk, Jens Nielsen, Lars K. Nielsen, Juan Nogales, Intawat Nookaew, Osbaldo Resendis, Bernhard Palsson, Jason A. Papin, Kiran Raosaheb Patil, Mark Poolman, Nathan D. Price, Anne Richelle, Isabel Rocha, Benjamin Sanchez, Peter Schaap, Rahuman S. Malik Sheriff, Saeed Shoaie, Nikolaus Sonnenschein, Bas Teusink, Paulo Vilaca, Jon Olav Vik, Judith A. Wodke, Joana C. Xavier, Qianqian Yuan, Maksim Zakhartsev, Cheng Zhang
Nat Biotechnol. 2020;38(3):272-276 (publication). doi:10.1038/s41587-020-0446-y. pmid:32123384
Reconstructing metabolic reaction networks enables the development of testable hypotheses of an organism’s metabolism under different conditions. State-of-the-art genome-scale metabolic models (GEMs) can include thousands of metabolites and reactions that are assigned to subcellular locations. Gene–protein–reaction (GPR) rules and annotations using database information can add meta-information to GEMs. GEMs with metadata can be built using standard reconstruction protocols, and guidelines have been put in place for tracking provenance and enabling interoperability, but a standardized means of quality control for GEMs is lacking. Here we report a community effort to develop a test suite named MEMOTE (for metabolic model tests) to assess GEM quality.
|
|
| 2020 |



|
Executable Simulation Model of the Liver
Matthias König
bioRxiv 2020.01.04.894873 (preprint). doi:10.1101/2020.01.04.894873
To address the issue of reproducibility in computational modeling we developed the concept of an executable simulation model (EXSIMO). An EXSIMO combines model, data and code with the execution environment to run the computational analysis in an automated manner using tools from software engineering. Key components are i) models, data and code for the computational analysis; ii) tests for models, data and code; and iii) an automation layer to run tests and execute the analysis. An EXSIMO combines version control, model, data, units, annotations, analysis, reports, execution environment, testing, continuous integration and release. We applied the concept to perform a replication study of a computational analysis of hepatic glucose metabolism in the liver. The corresponding EXSIMO is available from https://github.com/matthiaskoenig/exsimo
|
|
| 2019 |

|
Model Order Reduction (MOR) of Function-Perfusion-Growth Simulation in the Human Fatty Liver via Artificial Neural Network (ANN)
Lambers L., Ricken T., and König M.
Proceedings in Applied Mathematics & Mechanics, PAMM, 2019 (proceeding). doi:10.1002/pamm.201900429
Numerical modeling of biological systems has become an important assistance for understanding and predicting hepatic diseases like non-alcoholic fatty liver disease (NAFLD) or the detoxification of drugs and toxines by the liver. We developed a model for the simulation of hepatic function-perfusion processes using a multiscale and multiphase approach. Here, the liver lobules are described using a homogenization approach with a coupled set of partial differential equations (PDE) based on the Theory of Porous Media (TPM) to describe the coupled blood transport and tissue deformation. For the description of metabolic processes on cellular scale ordinary differential equations (ODE) are used. For many practical and clinical applications, e.g. optimization procedures or uncertainty quantification, a fast but reliable computation is required. Thus, we use a non-linear model order reduction (MOR) based on an artificial neural network (ANN) for the prediction of simulation results. The practicability of this approach is shown in a comparison between the high fidelity numerical simulation of a NAFLD and the predicted results by the ANN.
|
|
| 2019 |


|
Specifications of Standards in Systems and Synthetic Biology: Status and Developments in 2019
F Schreiber, G Bader, P Gleeson, M Golebiewski, M Hucka, SM Keating, M König, C Myers, D Nickerson, B Sommer, D Waltemath
J Integr Bioinform. 2019 Jul 13;16(2) (publication). doi:10.1515/jib-2019-0035. pmid:31301675
This special issue of the Journal of Integrative Bioinformatics presents an overview of COMBINE standards and their latest specifications. The standards cover representation formats for computational modeling in synthetic and systems biology and include BioPAX, CellML, NeuroML, SBML, SBGN, SBOL and SED-ML. The articles in this issue contain updated specifications of SBGN Process Description Level 1 Version 2, SBML Level 3 Core Version 2 Release 2, SBOL Version 2.3.0, and SBOL Visual Version 2.1.
|
|
| 2019 |



|
The Systems Biology Markup Language (SBML): Language Specification for Level 3 Version 2 Core
M. Hucka, F. Bergmann, C. Chaouiya, A. Dräger, S. Hoops, S. Keating, M. König, N Le Novere, C. Myers, B. Olivier, S. Sahle, J. Schaff, R. Sheriff, L. Smith, D. Waltemath, D. Wilkinson, F. Zhang
J Integr Bioinform. 2019 Jun 20;16(2); (publication). doi:10.1515/jib-2019-0021. pmid:31219795
Computational models can help researchers to interpret data, understand biological functions, and make quantitative predictions. The Systems Biology Markup Language (SBML) is a file format for representing computational models in a declarative form that different software systems can exchange. SBML is oriented towards describing biological processes of the sort common in research on a number of topics, including metabolic pathways, cell signaling pathways, and many others. By supporting SBML as an input/output format, different tools can all operate on an identical representation of a model, removing opportunities for translation errors and assuring a common starting point for analyses and simulations. This document provides the specification for Release 2 of Version 2 of SBML Level 3 Core. The specification defines the data structures prescribed by SBML as well as their encoding in XML, the eXtensible Markup Language. Release 2 corrects some errors and clarifies some ambiguities discovered in Release 1. This specification also defines validation rules that determine the validity of an SBML document, and provides many examples of models in SBML form. Other materials and software are available from the SBML project website at http://sbml.org/.
Keywords:
Representation,
Standards,
Systems Biology Markup Language,
Visualization,
|
|
| 2018 |



|
Tellurium: An Extensible Python-based Modeling Environment for Systems and Synthetic Biology
Choi K., Medley K., König M., Stocking K., Smith L., Gu S., and Sauro HM.
Biosystems. 2018 Jul 24. pii: S0303-2647(18)30125-4 (publication). doi:10.1016/j.biosystems.2018.07.006. pmid:30053414
Here we present Tellurium, a Python-based environment for model building, simulation, and analysis that facilitates reproducibility of models in systems and synthetic biology. Tellurium is a modular, cross-platform, and open-source simulation environment composed of multiple libraries, plugins, and specialized modules and methods. Tellurium is a self-contained modeling platform which comes with a fully configured Python distribution. Two interfaces are provided, one based on the Spyder IDE which has an accessible user interface akin to MATLAB and a second based on the Jupyter Notebook, which is a format that contains live code, equations, visualizations and narrative text. Tellurium uses libRoadRunner as the default SBML simulation engine which supports deterministic simulations, stochastic simulations, and steady-state analyses. Tellurium also includes Antimony, a human-readable model definition language which can be converted to and from SBML. Other standard Python scientific libraries such as NumPy, SciPy, and matplotlib are included by default. Additionally, we include several user-friendly plugins and advanced modules for a wide-variety of applications, ranging from complex algorithms for bifurcation analysis to multidimensional parameter scanning. By combining multiple libraries, plugins, and modules into a single package, Tellurium provides a unified but extensible solution for biological modeling and analysis for both novices and experts.
|
|
| 2018 |

|
HEPATOKIN1: A Biochemistry-Based Model of Liver Metabolism Suited for Applications in Medicine and Pharmacology
Berndt N., Bulik S., Wallach I., Wünsch T., König M., Stockmann M., Meierhofer M., Holzhütter HG.
Nat Commun. 2018 Jun 19;9(1):2386. (publication). doi:10.1038/s41467-018-04720-9. pmid:29921957
The most reliable and predictive type of metabolic models are biochemistry-based kinetic models resting on the principles of chemical kinetics and including all relevant molecular details of enzyme regulation. Here we present such a model with yet unequalled complexity encompassing the major pathways of hepatic energy, carbohydrate, lipid, and nitrogen metabolism and including the regulation of enzyme activities by their reactants, allosteric effectors and hormone-dependent reversible enzyme phosphorylation. We illustrate the utility of the model for basic research and applications in medicine and pharmacology by simulating diurnal variations of the metabolic state of the liver under normal conditions and at various perturbations caused by nutritional challenges (alcohol), drugs (valproate) and inherited enzyme disorders (galactosemia). In particular, the model can be used to quantify alterations in the metabolic functions of different liver cells (normal hepatocytes versus adenoma and hepatocellular carcinoma) by exploiting relative changes in the abundance of metabolic enzymes for the re-scaling of maximal enzyme activities (Vmax values).
|
|
| 2018 |



|
Simulation Experiment Description Markup Language (SED-ML) Level 1 Version 3 (L1V3)
Bergmann, FT., Cooper, J., König, M.Δ, Moraru, I., Nickerson, D., Le Novère N., Olivier, BG., Sahle, S., Smith, L., Waltemath, D. Δ corresponding author
J Integr Bioinform. 2018 Mar 19;15(1) (publication). doi:10.1515/jib-2017-0086. pmid:29550789
The creation of computational simulation experiments to inform modern biological research poses challenges to reproduce, annotate, archive, and share such experiments. Efforts such as SBML or CellML standardize the formal representation of computational models in various areas of biology. The Simulation Experiment Description Markup Language (SED-ML) describes what procedures the models are subjected to, and the details of those procedures. These standards, together with further COMBINE standards, describe models sufficiently well for the reproduction of simulation studies among users and software tools. The Simulation Experiment Description Markup Language (SED-ML) is an XML-based format that encodes, for a given simulation experiment, i) which models to use; ii) which modifications to apply to models before simulation; iii) which simulation procedures to run on each model; iv) how to post-process the data; and v) how these results should be plotted and reported. SED-ML Level 1 Version 1 (L1V1) implemented support for the encoding of basic time course simulations. SED-ML L1V2 added support for more complex types of simulations, specifically repeated tasks and chained simulation procedures. SED-ML L1V3 extends L1V2 by means to describe which datasets and subsets thereof to use within a simulation experiment.
|
|
| 2018 |



|
Harmonizing semantic annotations for computational models in biology
Neal, M.; König, M.; Nickerson, D.; Misirli, G.; Kalbasi, R.; Dräger, A.; Atalag, K.; Chelliah, V.; Cooling, M.; Cook, D.; Crook, S.; de Alba, M.; Friedman, S.; Garny, A.; Gennari, J.; Gleeson, P.; Golebiewski, M.; Hucka, M.; Juty, N.; Novère, N. L.; Myers, C.; Olivier, B.; Sauro, H.; Scharm, M.; Snoep, J.; Touré, V.; Wipat, A.; Wolkenhauer, O. & Waltemath, D.
Brief Bioinform. 2019 Mar 22;20(2):540-550 (publication). doi:10.1093/bib/bby087. pmid:30462164
Life science researchers use computational models to articulate and test hypotheses about the behavior of biological systems. Semantic annotation is a critical component for enhancing the interoperability and reusability of such models as well as for the integration of the data needed for model parameterization and validation. Encoded as machine-readable links to knowledge resource terms, semantic annotations describe the computational or biological meaning of what models and data represent. These annotations help researchers find and repurpose models, accelerate model composition, and enable knowledge integration across model repositories and experimental data stores. However, realizing the potential benefits of semantic annotation requires the development of model annotation standards that adhere to a community-based annotation protocol. Without such standards, tool developers must account for a variety of annotation formats and approaches, a situation that can become prohibitively cumbersome and which can defeat the purpose of linking model elements to controlled knowledge resource terms. Currently, no consensus protocol for semantic annotation exists among the larger biological modeling community. Here, we report on the landscape of current semantic annotation practices among the COmputational Modeling in BIology NEtwork (COMBINE) community and provide a set of recommendations for building a consensus approach to semantic annotation.
|
|
| 2018 |



|
Tellurium Notebooks — An Environment for Reproducible Dynamical Modeling in Systems Biology.
Medley K., Choi K., König M., Smith L., Gu S., Hellerstein J., Sealfon S., and Sauro HM.
PLoS Comput Biol. 2018 Jun 15;14(6):e1006220 (publication). doi:10.1371/journal.pcbi.1006220. pmid:29906293
The considerable difficulty encountered in reproducing the results of published dynamical models limits validation, exploration and reuse of this increasingly large biomedical research resource. To address this problem, we have developed Tellurium Notebook, a software system that facilitates building reproducible dynamical models and reusing models by 1) supporting the COMBINE archive format during model development for capturing model information in an exchangeable format and 2) enabling users to easily simulate and edit public COMBINE-compliant models from public repositories to facilitate studying model dynamics, variants and test cases. Tellurium Notebook, a Python-based Jupyter-like environment, is designed to seamlessly inter-operate with these community standards by automating conversion between COMBINE standards formulations and corresponding in-line, human-readable representations. Thus, Tellurium brings to systems biology the strategy used by other literate notebook systems such as Mathematica. These capabilities allow users to edit every aspect of the standards-compliant models and simulations, run the simulations in-line, and re-export to standard formats. We provide several use cases illustrating the advantages of our approach and how it allows development and reuse of models without requiring technical knowledge of standards. Adoption of Tellurium should accelerate model development, reproducibility and reuse.
|
|
| 2017 |

|
Computational Modeling in Liver Surgery
Christ, B.Δ, Dahmen, U.Δ, Herrmann, K.H.Δ, König, M.Δ, Reichenbach, J.RΔ., Ricken, T.Δ, Schleicher, J.Δ, Ole Schwen, L.Δ, Vlaic, S.Δ and Waschinsky, N.Δ (Δ equal contribution)
Front Physiol. 2017 Nov 14;8:906 (review). doi:10.3389/fphys.2017.00906. pmid:29249974
The need for extended liver resection is increasing due to the growing incidence of liver tumors in aging societies. Individualized surgical planning is the key for identifying the optimal resection strategy and to minimize the risk of postoperative liver failure and tumor recurrence. Current computational tools provide virtual planning of liver resection by taking into account the spatial relationship between the tumor and the hepatic vascular trees, as well as the size of the future liver remnant. However, size and function of the liver are not necessarily equivalent. Hence, determining the future liver volume might misestimate the future liver function, especially in cases of hepatic comorbidities such as hepatic steatosis. A systems medicine approach could be applied, including biological, medical, and surgical aspects, by integrating all available anatomical and functional information of the individual patient. Such an approach holds promise for better prediction of postoperative liver function and hence improved risk assessment. This review provides an overview of mathematical models related to the liver and its function and explores their potential relevance for computational liver surgery. We first summarize key facts of hepatic anatomy, physiology, and pathology relevant for hepatic surgery, followed by a description of the computational tools currently used in liver surgical planning. Then we present selected state-of-the-art computational liver models potentially useful to support liver surgery. Finally, we discuss the main challenges that will need to be addressed when developing advanced computational planning tools in the context of liver surgery.
Keywords:
Liver resection,
function prediction,
liver metabolism,
liver regeneration,
liver surgical planning,
multi-scale modeling,
risk assessment,
systems medicine,
|
|
| 2016 |

|
Challenges and opportunities for system biology standards and tools in medical research
König M.Δ, Oellrich A.Δ, Waltemath D.Δ, Dobson R., Hubbard T., and Wolkenhauer O. (Δ equal contribution)
ODLS, p1-6, 2016 (proceeding)
Kinetic models are increasingly relevant in medical research. In systems biology, more than 10 years of experience with the development of standards and tools to construct and analyse kinetic models exists. This has supported the sharing of kinetic models, increased their reuse, and thereby has helped to reproduce and validate scientific results. Given the expertise and the existing infrastructure, it seems natural to consider the application and development of standards and tools to meet the requirements of medical scientists. In this paper, we discuss challenges and opportunities for standards and tools from systems biology in medical research, and we put forward criteria for the safe use of simulations. We conclude that standards, tools and infrastructure need to be extended to ensure the quality, reliability and safety required when working with medical and patient data. This will foster the adaptation of modelling in the clinic, providing tools for improved diagnosis, prognosis and therapy.
|
|
| 2016 |



|
cy3sabiork: A Cytoscape app for visualizing kinetic data from SABIO-RK
König M.
[version 1; referees: awaiting peer review]. F1000Research 2016, 5:1736 (preprint). doi:10.12688/f1000research.9211.1
Kinetic data of biochemical reactions are essential for the creation of kinetic models of biochemical networks. One of the main resources of such information is SABIO-RK, a curated database for kinetic data of biochemical reactions and their related information. Despite the importance for computational modelling there has been no simple solution to visualize the kinetic data from SABIO-RK. In this work, I present cy3sabiork, an app for querying and visualization of kinetic data from SABIO-RK in Cytoscape. The kinetic information is accessible via a combination of graph structure and annotations of nodes, with provided information consisting of: (I) reaction details, enzyme and organism; (II) kinetic law, formula, parameters; (III) experimental conditions; (IV) publication; (V) additional annotations. cy3sabiork creates an intuitive visualization of kinetic entries in form of a species-reaction-kinetics graph, which reflects the reaction-centered approach of SABIO-RK. Kinetic entries can be imported in SBML format from either the SABIO-RK web interface or via web service queries. The app allows for easy comparison of kinetic data, visual inspection of the elements involved in the kinetic record and simple access to the annotation information of the kinetic record. I applied cy3sabiork in the computational modelling of galactose metabolism in the human liver.
|
|
| 2016 |



|
Toward community standards and software for whole-cell modeling
Wholecell Consortium (König M.)
IEEE Trans Biomed Eng. 2016 Jun 10. (publication). doi:10.1109/TBME.2016.2560762. pmid:27305665
Objective: Whole-cell (WC) modeling is a promising tool for biological research, bioengineering, and medicine. However, substantial work remains to create accurate comprehensive models of complex cells. Methods: We organized the 2015 Whole-Cell Modeling Summer School to teach WC modeling and evaluate the need for new WC modeling standards and software by recoding a recently published WC model in the Systems Biology Markup Language. Results: Our analysis revealed several challenges to representing WC models using the current standards. Conclusion: We, therefore, propose several new WC modeling standards, software, and databases. Significance: We anticipate that these new standards and software will enable more comprehensive models.
|
|
| 2015 |



|
Pathobiochemical signatures of cholestatic liver disease in bile duct ligated mice
Kerstin A.Δ, König M.Δ, Hoppe A., Thomas M., Müller I., Ebert M., Weng H., Holzhütter HG., Zanger UM., Bode J., Vollmar B. and Dooley S. (Δ equal contribution)
BMC Syst Biol. 2015 Nov 20;9(1):83. (publication). doi:10.1186/s12918-015-0229-0. pmid:26589287
Background: Disrupted bile secretion leads to liver damage characterized by inflammation, fibrosis, eventually cirrhosis, and hepatocellular cancer. As obstructive cholestasis often progresses insidiously, markers for the diagnosis and staging of the disease are urgently needed. To this end, we compiled a comprehensive data set of serum markers, histological parameters and transcript profiles at 8 time points of disease progression after bile duct ligation (BDL) in mice, aiming at identifying a set of parameters that could be used as robust biomarkers for transition of different disease progression phases. Results: Statistical analysis of the more than 6,000 data points revealed distinct temporal phases of disease. Time course correlation analysis of biochemical, histochemical and mRNA transcript parameters (=factors) defined 6 clusters for different phases of disease progression. The number of CTGF-positive cells provided the most reliable overall measure for disease progression at histological level, bilirubin at biochemical level, and metalloproteinase inhibitor 1 (Timp1) at transcript level. Prominent molecular events exhibited by strong transcript peaks are found for the transcriptional regulator Nr0b2 (Shp) and 1,25-dihydroxyvitamin D(3) 24-hydroxylase (Cyp24a1) at 6 h. Based on these clusters, we constructed a decision tree of factor combinations potentially useful as markers for different time intervals of disease progression. Best prediction for onset of disease is achieved by fibronectin (Fn1), for early disease phase by Cytochrome P450 1A2 (Cyp1a2), passage to perpetuation phase by collagen1α-1 (Col1a1), and transition to the progression phase by interleukin 17-a (Il17a), with early and late progression separated by Col1a1. Notably, these predictions remained stable even for randomly chosen small sub-sets of factors selected from the clusters. Conclusion: Our detailed time-resolved explorative study of liver homogenates following BDL revealed a well-coordinated response, resulting in disease phase dependent parameter modulations at morphological, biochemical, metabolic and gene expression levels. Interestingly, a small set of selected parameters can be used as diagnostic markers to predict disease stages in mice with cholestatic liver disease.
|
|
| 2015 |
|
On the Influence of Growth in Perfusion Dependent Biological Systems - at the Example of the Human Liver.
Werner D., Ricken T., Dahmen U., Dirsch O., Holzhütter HG., König M.
PAMM 15 (1), 119-120, 2015 (proceeding). doi:10.1002/pamm.201510050
Biochemical reactions in the liver are a complex non-linear and time depending coupled function-perfusion-mechanisms. The metabolism of the liver, e.g. nutrient storage, is directly coupled to its blood perfusion. Growth phenomena, such as accumulation of fat, have a strong impact onto micro-perfusion within the liver. Here, we present the results of a computational model that describes selected coupled functionalities. The metabolic processes take place in the liver cells, the hepatocytes, which are arranged in a delicate system of capillaries, so called sinusoids. Nutrients, oxygen, and other components of the blood are transported through the sinusoids. Due to this highly complex inner structure of the lobules it is impracticable to give a microscopic geometrical description in a continuum mechanical manner. Therefore, a homogenization scheme based on the theory of porous media (TPM) is used;. In case of the presented liver model we consider a porous solid body, that consists of tissue and fat, and a fluid representing blood. In each phase several miscible components are considered. For the microscopic cell level use has been made of an embedded set of ordinary differential equations (ODE) that mimics a simplified metabolism, e.g. glucose utilization and production;. In case of growth of the fat phase we follow a phenomenological approach. Numerical examples for this coupled process are presented and discussed.
|
|
| 2015 |



|
LibRoadRunner: a high performance SBML simulation and analysis library.
Somogyi ET., Bouteiller JM., Glazier JA., König M., Medley JK., Swat MH and Sauro HM.
Bioinformatics. 2015 Oct 15;31(20):3315-21 (publication). doi:10.1093/bioinformatics/btv363. pmid:26085503
Motivation: This article presents libRoadRunner, an extensible, high-performance, cross-platform, open-source software library for the simulation and analysis of models expressed using Systems Biology Markup Language (SBML). SBML is the most widely used standard for representing dynamic networks, especially biochemical networks. libRoadRunner is fast enough to support large-scale problems such as tissue models, studies that require large numbers of repeated runs and interactive simulations. Results: libRoadRunner is a self-contained library, able to run both as a component inside other tools via its C++ and C bindings, and interactively through its Python interface. Its Python Application Programming Interface (API) is similar to the APIs of MATLAB and SciPy, making it fast and easy to learn. libRoadRunner uses a custom Just-In-Time (JIT) compiler built on the widely used LLVM JIT compiler framework. It compiles SBML-specified models directly into native machine code for a variety of processors, making it appropriate for solving extremely large models or repeated runs. libRoadRunner is flexible, supporting the bulk of the SBML specification (except for delay and non-linear algebraic equations) including several SBML extensions (composition and distributions). It offers multiple deterministic and stochastic integrators, as well as tools for steady-state analysis, stability analysis and structural analysis of the stoichiometric matrix. Availability and implementation: libRoadRunner binary distributions are available for Mac OS X, Linux and Windows. The library is licensed under Apache License Version 2.0. libRoadRunner is also available for ARM-based computers such as the Raspberry Pi. http://www.libroadrunner.org provides online documentation, full build instructions, binaries and a git source repository.
|
|
| 2014 |

|
Modeling function-perfusion behavior in liver lobules including tissue, blood, glucose, lactate and glycogen by use of a coupled two-scale PDE-ODE approach.
Ricken T., Werner D., Holzhütter HG., König M., Dahmen U., Dirsch O.
Biomech Model Mechanobiol. 2015 Jun;14(3):515-36 (publication). doi:10.1007/s10237-014-0619-z. pmid:25236798
This study focuses on a two-scale, continuum multicomponent model for the description of blood perfusion and cell metabolism in the liver. The model accounts for a spatial and time depending hydro-diffusion-advection-reaction description. We consider a solid-phase (tissue) containing glycogen and a fluid-phase (blood) containing glucose as well as lactate. The five-component model is enhanced by a two-scale approach including a macroscale (sinusoidal level) and a microscale (cell level). The perfusion on the macroscale within the lobules is described by a homogenized multiphasic approach based on the theory of porous media (mixture theory combined with the concept of volume fraction). On macro level, we recall the basic mixture model, the governing equations as well as the constitutive framework including the solid (tissue) stress, blood pressure and solutes chemical potential. In view of the transport phenomena, we discuss the blood flow including transverse isotropic permeability, as well as the transport of solute concentrations including diffusion and advection. The continuum multicomponent model on the macroscale finally leads to a coupled system of partial differential equations (PDE). In contrast, the hepatic metabolism on the microscale (cell level) was modeled via a coupled system of ordinary differential equations (ODE). Again, we recall the constitutive relations for cell metabolism level. A finite element implementation of this framework is used to provide an illustrative example, describing the spatial and time-depending perfusion-metabolism processes in liver lobules that integrates perfusion and metabolism of the liver.
|
|
| 2014 |

|
Homeostasis of blood glucose - Computer simulations of central liver functions.
König M. and Holzhütter HG.
systembiologie.de 2014; 8:p.53-57 (proceeding)
The liver performs a multitude of important physiological functions such as synthesising plasma proteins, producing bile and hormones, detoxifying toxic compounds and precisely regulating the concentration of various substances in the blood (homeostasis). In particular, the liver plays a crucial role in the regulation of blood glucose levels. Modelling hepatic glucose metabolism helps to understand central liver functions and basic regulation mechanisms and thereby provides insights in the role of the liver in systemic diseases such as diabetes.
|
|
| 2013 |

|
Metabolic Gradients as Key Regulators in Zonation of Tumor Energy Metabolism: A Tissue-scale Model Based Study.
König M., Holzhütter HG., Berndt N.
Biotechnol J. 2013 Sep;8(9):1058-69. (publication). doi:10.1002/biot.201200393. pmid:23589477
Characteristics of many tumor types are the reprogramming of metabolism and the occurrence of regional hypoxia. In this work, we investigated the hypothesis that metabolic reprogramming in combination with metabolic zonation of cellular energy metabolism are important factors in promotion of the growth capacity of solid tumors. A tissue-scale model of the two main ATP delivering pathways, glycolysis (GLY) and oxidative phosphorylation (OXP), was used to simulate the energy metabolism within solid tumors under various metabolic strategies. Remarkably, despite the high diversity in the usage of glucose, lactate and oxygen in various spatial regions, the tumor as a whole clearly displays the hallmark of the so-called Warburg effect, i.e. a high rate of glucose consumption and lactate production in the presence of sufficiently high levels of oxygen. Our simulations suggest that an increase in GLY capacity and concomitant decrease in OXP capacity from the periphery towards the center of the tumor improves the availability of oxygen to pericentral tumor cells. The found relationship between the regional oxygen level and the relative share of GLY and OXP capacities supports the view that metabolite availability functions as key regulator of tumor energy metabolism.
|
|
| 2012 |

|
Kinetic Modeling of Human Hepatic Glucose Metabolism in T2DM Predicts Higher Risk of Hypoglycemic Events in Rigorous Insulin Therapy
König M. and Holzhütter HG.
J Biol Chem. 2012 Oct 26;287(44):36978-89 (publication). doi:10.1074/jbc.M112.382069. pmid:22977253
A major problem in the insulin therapy of patients with diabetes type 2 (T2DM) is the increased occurrence of hypoglycemic events which, if left untreated, may cause confusion or fainting and in severe cases seizures, coma, and even death. To elucidate the potential contribution of the liver to hypoglycemia in T2DM we applied a detailed kinetic model of human hepatic glucose metabolism to simulate changes in glycolysis, gluconeogenesis, and glycogen metabolism induced by deviations of the hormones insulin, glucagon, and epinephrine from their normal plasma profiles. Our simulations reveal in line with experimental and clinical data from a multitude of studies in T2DM, (i) significant changes in the relative contribution of glycolysis, gluconeogenesis, and glycogen metabolism to hepatic glucose production and hepatic glucose utilization; (ii) decreased postprandial glycogen storage as well as increased glycogen depletion in overnight fasting and short term fasting; and (iii) a shift of the set point defining the switch between hepatic glucose production and hepatic glucose utilization to elevated plasma glucose levels, respectively, in T2DM relative to normal, healthy subjects. Intriguingly, our model simulations predict a restricted gluconeogenic response of the liver under impaired hormonal signals observed in T2DM, resulting in an increased risk of hypoglycemia. The inability of hepatic glucose metabolism to effectively counterbalance a decline of the blood glucose level becomes even more pronounced in case of tightly controlled insulin treatment. Given this Janus face mode of action of insulin, our model simulations underline the great potential that normalization of the plasma glucagon profile may have for the treatment of T2DM.
|
|
| 2012 |



|
CySBML: a Cytoscape plugin for SBML
König M., Dräger A. and Holzhütter HG.
Bioinformatics. 2012 Sep 15;28(18):2402-3 (publication). doi:10.1093/bioinformatics/bts432. pmid:22772946
Summary: CySBML is a plugin designed to work with Systems Biology Markup Language (SBML) in Cytoscape having the following features: SBML import, support of the SBML layout and qualitative model packages, navigation in network layouts based on SBML structure, access to MIRIAM and SBO-based annotations and SBML validation. CySBML includes an importer for BioModels to load SBML from standard repositories. Availability and implementation: Freely available for non-commercial purposes through the Cytoscape plugin manager or for download at http://sourceforge.net/projects/cysbml/. Contact: cysbml-team@lists.sourceforge.net. Supplementary information: Tutorial, usage guide, installation instructions and additional figures are available for download at http://www.charite.de/sysbio/people/koenig/software/cysbml/.
|
|
| 2012 |

|
Quantifying the Contribution of the Liver to the Homeostasis of Plasma Glucose: A Detailed Kinetic Model of Hepatic Glucose Metabolism Integrated with the Hormonal Control by Insulin, Glucagon and Epinephrine
König M., Bulik S. and Holzhütter HG.
PLoS Comput Biol. 2012 Jun;8(6):e1002577. Epub 2012 Jun 21. (publication). doi:10.1371/journal.pcbi.1002577. pmid:22761565
Despite the crucial role of the liver in glucose homeostasis, a detailed mathematical model of human hepatic glucose metabolism is lacking so far. Here we present a detailed kinetic model of glycolysis, gluconeogenesis and glycogen metabolism in human hepatocytes integrated with the hormonal control of these pathways by insulin, glucagon and epinephrine. Model simulations are in good agreement with experimental data on (i) the quantitative contributions of glycolysis, gluconeogenesis, and glycogen metabolism to hepatic glucose production and hepatic glucose utilization under varying physiological states. (ii) the time courses of postprandial glycogen storage as well as glycogen depletion in overnight fasting and short term fasting (iii) the switch from net hepatic glucose production under hypoglycemia to net hepatic glucose utilization under hyperglycemia essential for glucose homeostasis (iv) hormone perturbations of hepatic glucose metabolism. Response analysis reveals an extra high capacity of the liver to counteract changes of plasma glucose level below 5 mM (hypoglycemia) and above 7.5 mM (hyperglycemia). Our model may serve as an important module of a whole-body model of human glucose metabolism and as a valuable tool for understanding the role of the liver in glucose homeostasis under normal conditions and in diseases like diabetes or glycogen storage diseases.
|
|
| 2011 |

|
Enzymatic Features of the Glucose Metabolism in Tumor Cells.
Herling A, König M, Bulik S, Holzhütter HG.
FEBS J. 2011 Jul;278(14):2436-59 (review). doi:10.1111/j.1742-4658.2011.08174.x. pmid:21564549
Many tumor types exhibit an impaired Pasteur effect, i.e. despite the presence of oxygen, glucose is consumed at an extraordinarily high rate compared with the tissue from which they originate - the so-called "Warburg effect". Glucose has to serve as the source for a diverse array of cellular functions, including energy production, synthesis of nucleotides and lipids, membrane synthesis and generation of redox equivalents for antioxidative defense. Tumor cells acquire specific enzyme-regulatory mechanisms to direct the main flux of glucose carbons to those pathways most urgently required under challenging external conditions such as varying substrate availability, presence of anti-cancer drugs or different phases of the cell cycle. In this review we summarize the currently available information on tumor-specific expression, activity and kinetic properties of enzymes involved in the main pathways of glucose metabolism with due regard to the explanation of the regulatory basis and physiological significance of the Warburg effect. We conclude that, besides the expression level of the metabolic enzymes involved in the glucose metabolism of tumor cells, the unique tumor-specific pattern of isozymes and accompanying changes in the metabolic regulation below the translation level enable tumor cells to drain selfishly the blood glucose pool that non-transformed cells use as sparingly as possible.
|
|
| 2011 |



|
FluxViz - Cytoscape Plug-in for Vizualisation of Flux Distributions in Networks
König M. and Holzhütter HG.
Genome Informatics 2010, Vol.24, p.96-103 (proceeding). pmid:22081592
Motivation: Methods like FBA and kinetic modeling are widely used to calculate fluxes in metabolic networks. For the analysis and understanding of simulation results and experimentally measured fluxes visualization software within the network context is indispensable. Results: We present Flux Viz, an open-source Cytoscape plug-in for the visualization of flux distributions in molecular interaction networks. FluxViz supports (i) import of networks in a variety of formats (SBML, GML, XGMML, SIF, BioPAX, PSI-MI) (ii) import of flux distributions as CSV, Cytoscape attributes or VAL files (iii) limitation of views to flux carrying reactions (flux subnetwork) or network attributes like localization (iv) export of generated views (SVG, EPS, PDF, BMP, PNG). Though FluxViz was primarily developed as tool for the visualization of fluxes in metabolic networks and the analysis of simulation results from FASIMU, a flexible software for batch flux-balance computation in large metabolic networks, it is not limited to biochemical reaction networks and FBA but can be applied to the visualization of arbitrary fluxes in arbitrary graphs. Availability: The platform-independent program is an open-source project, freely available at http://sourceforge.net/projects/fluxvizplugin/ under GNU public license, including manual, tutorial and examples.
|
|
| 2010 |

|
HepatoNet1: a comprehensive metabolic reconstruction of the human hepatocyte for the analysis of liver physiology.
Gille C, Bölling C, Hoppe A, Bulik S, Hoffmann S, Hübner K, Karlstädt A, Ganeshan R, König M, Rother K, Weidlich M, Behre J, Holzhütter HG.
Mol Syst Biol. 2010 Sep 7;6:411 (publication). doi:10.1038/msb.2010.62. pmid:20823849
We present HepatoNet1, the first reconstruction of a comprehensive metabolic network of the human hepatocyte that is shown to accomplish a large canon of known metabolic liver functions. The network comprises 777 metabolites in six intracellular and two extracellular compartments and 2539 reactions, including 1466 transport reactions. It is based on the manual evaluation of >1500 original scientific research publications to warrant a high-quality evidence-based model. The final network is the result of an iterative process of data compilation and rigorous computational testing of network functionality by means of constraint-based modeling techniques. Taking the hepatic detoxification of ammonia as an example, we show how the availability of nutrients and oxygen may modulate the interplay of various metabolic pathways to allow an efficient response of the liver to perturbations of the homeostasis of blood compounds.
|
|
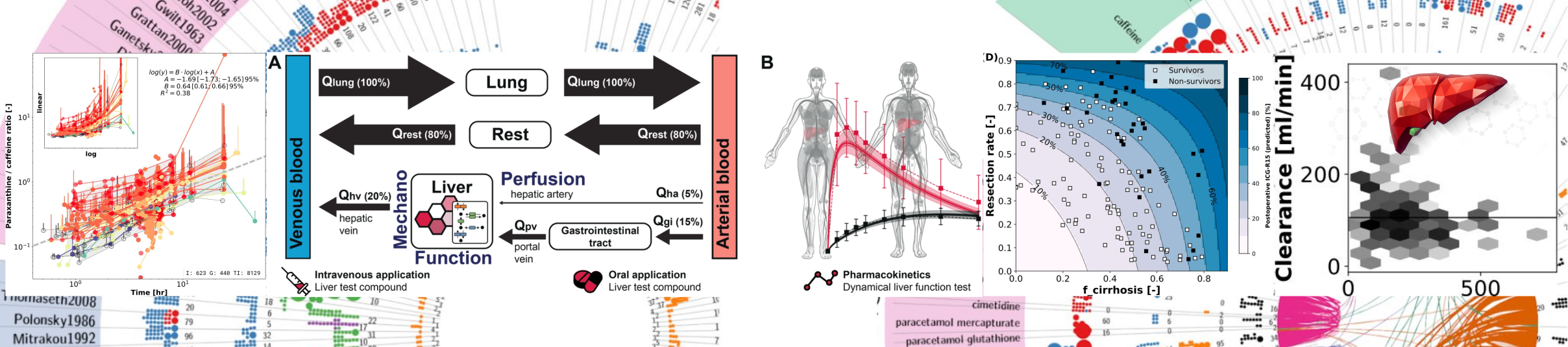













 We present PK-DB 2.0, an open, FAIR, and AI-ready pharmacokinetics and pharmacodynamics database providing curated PK/PD data from more than 700 clinical studies for pharmacometric modeling, machine learning, and digital twin development.
We present PK-DB 2.0, an open, FAIR, and AI-ready pharmacokinetics and pharmacodynamics database providing curated PK/PD data from more than 700 clinical studies for pharmacometric modeling, machine learning, and digital twin development.
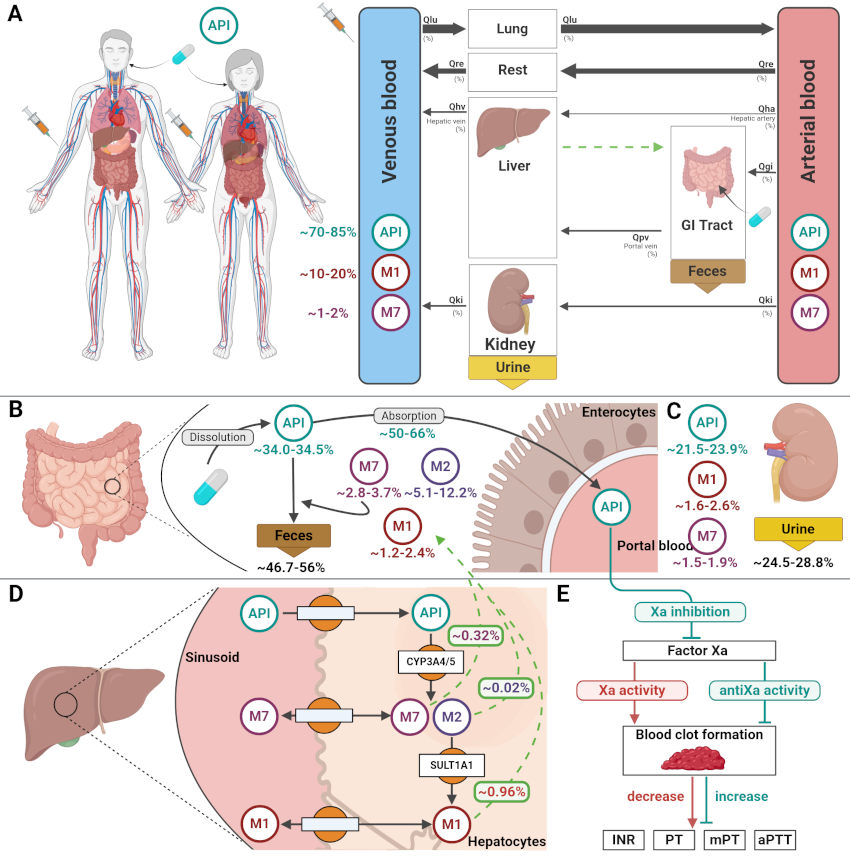 We present an open whole-body PBPK/PD digital twin of apixaban to investigate pharmacokinetic and pharmacodynamic variability in personalised anticoagulant therapy, with a focus on renal and hepatic impairment, food intake, body weight, metabolites, and coagulation-related endpoints.
We present an open whole-body PBPK/PD digital twin of apixaban to investigate pharmacokinetic and pharmacodynamic variability in personalised anticoagulant therapy, with a focus on renal and hepatic impairment, food intake, body weight, metabolites, and coagulation-related endpoints.
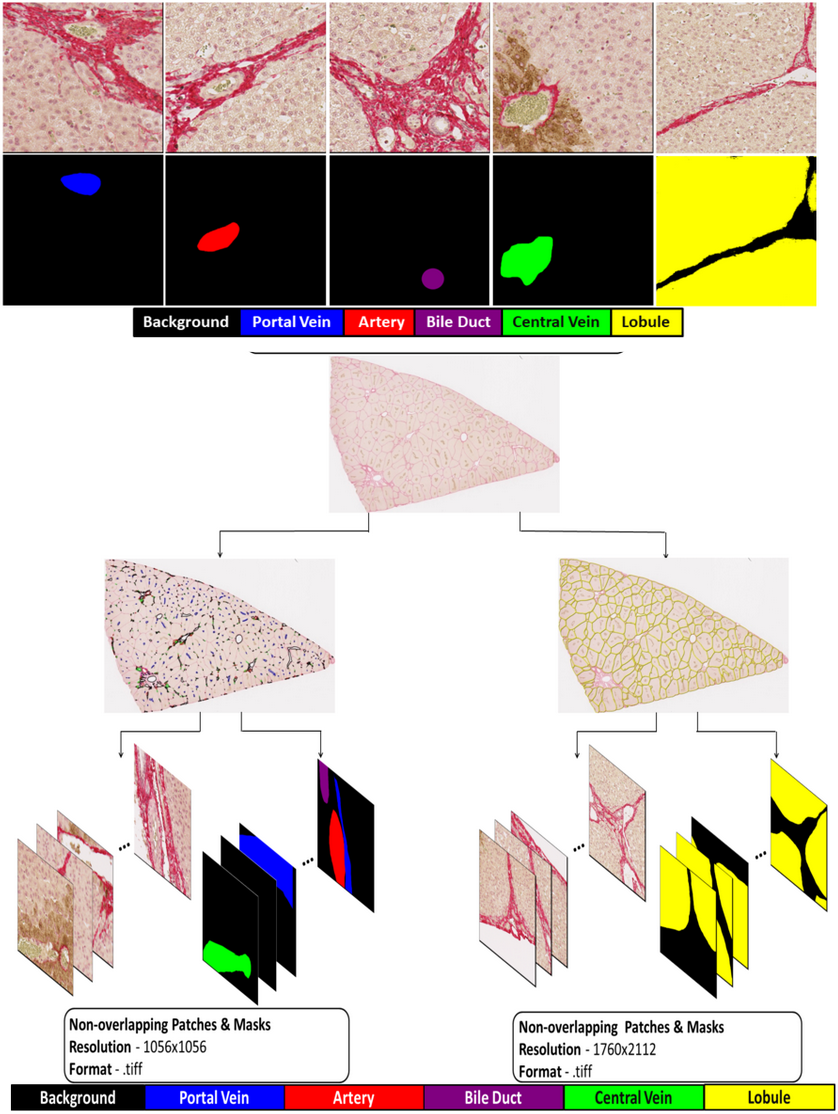 Our latest publication introduces a deep-learning pipeline for automated segmentation of hepatic vessels, bile ducts, and lobules in whole-slide liver histology images.
Our latest publication introduces a deep-learning pipeline for automated segmentation of hepatic vessels, bile ducts, and lobules in whole-slide liver histology images.
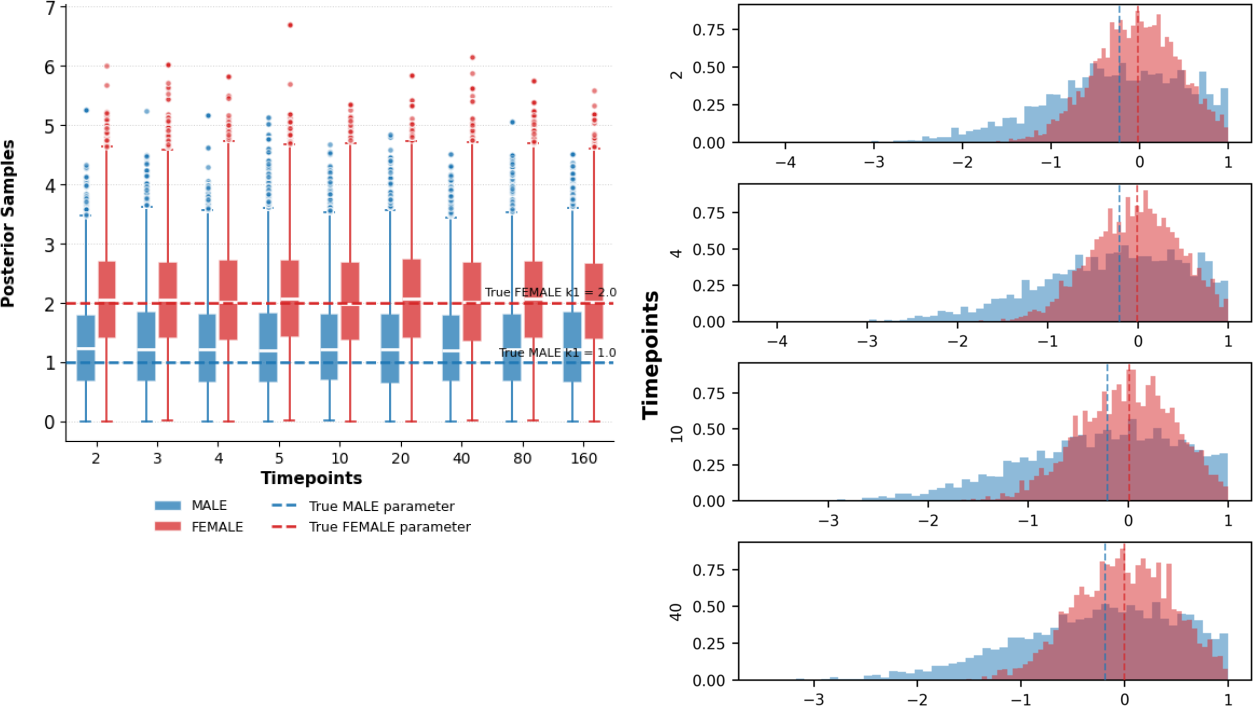 Antonio Alvarez submitted his Master thesis on reproducible Bayesian optimization and uncertainty quantification in pharmacokinetic models
Antonio Alvarez submitted his Master thesis on reproducible Bayesian optimization and uncertainty quantification in pharmacokinetic models
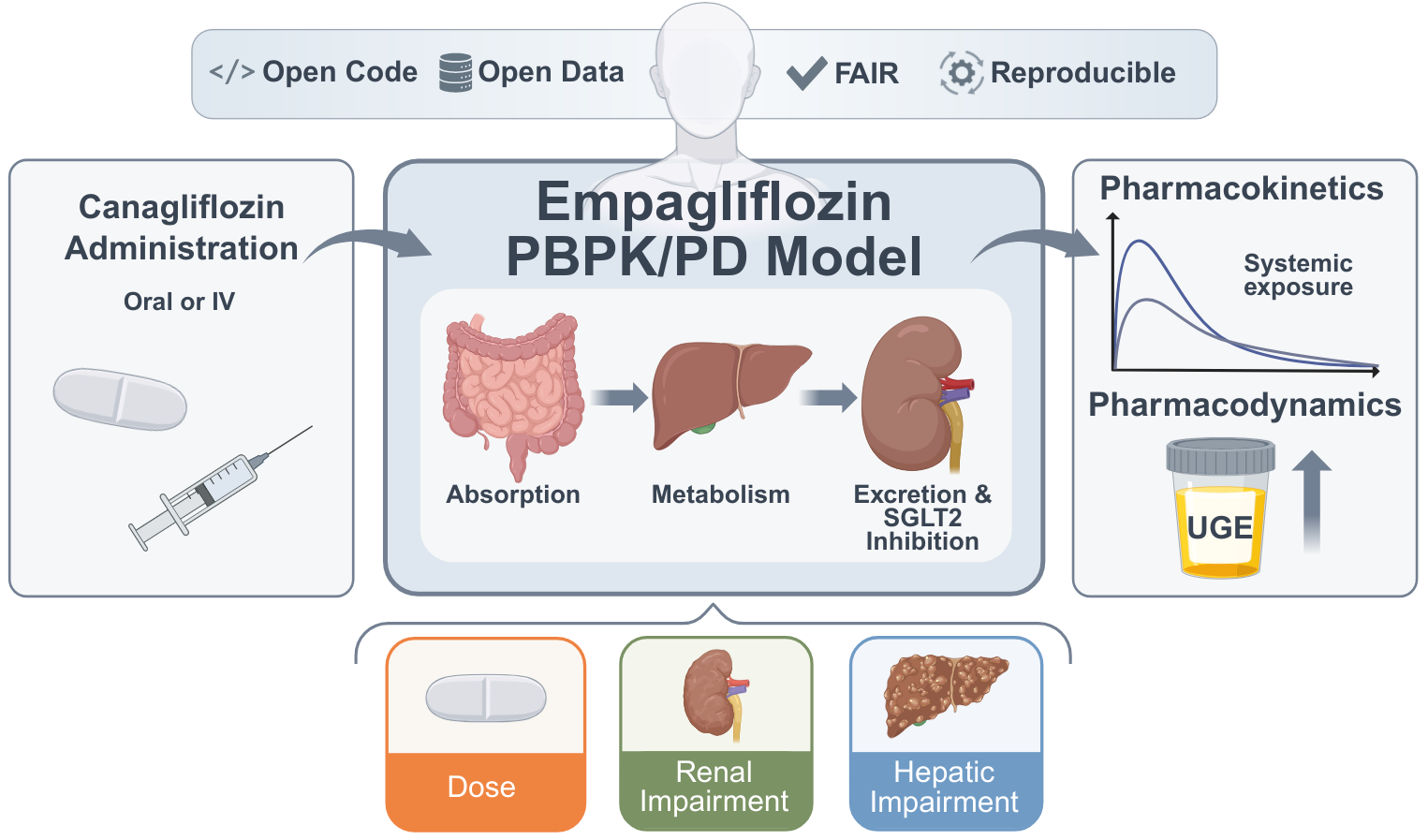 We present a whole-body PBPK/PD digital twin of empagliflozin to investigate pharmacokinetic and pharmacodynamic variability in type 2 diabetes mellitus, with a focus on renal and hepatic impairment, metabolite exposure, and dose-dependent effects.
We present a whole-body PBPK/PD digital twin of empagliflozin to investigate pharmacokinetic and pharmacodynamic variability in type 2 diabetes mellitus, with a focus on renal and hepatic impairment, metabolite exposure, and dose-dependent effects.
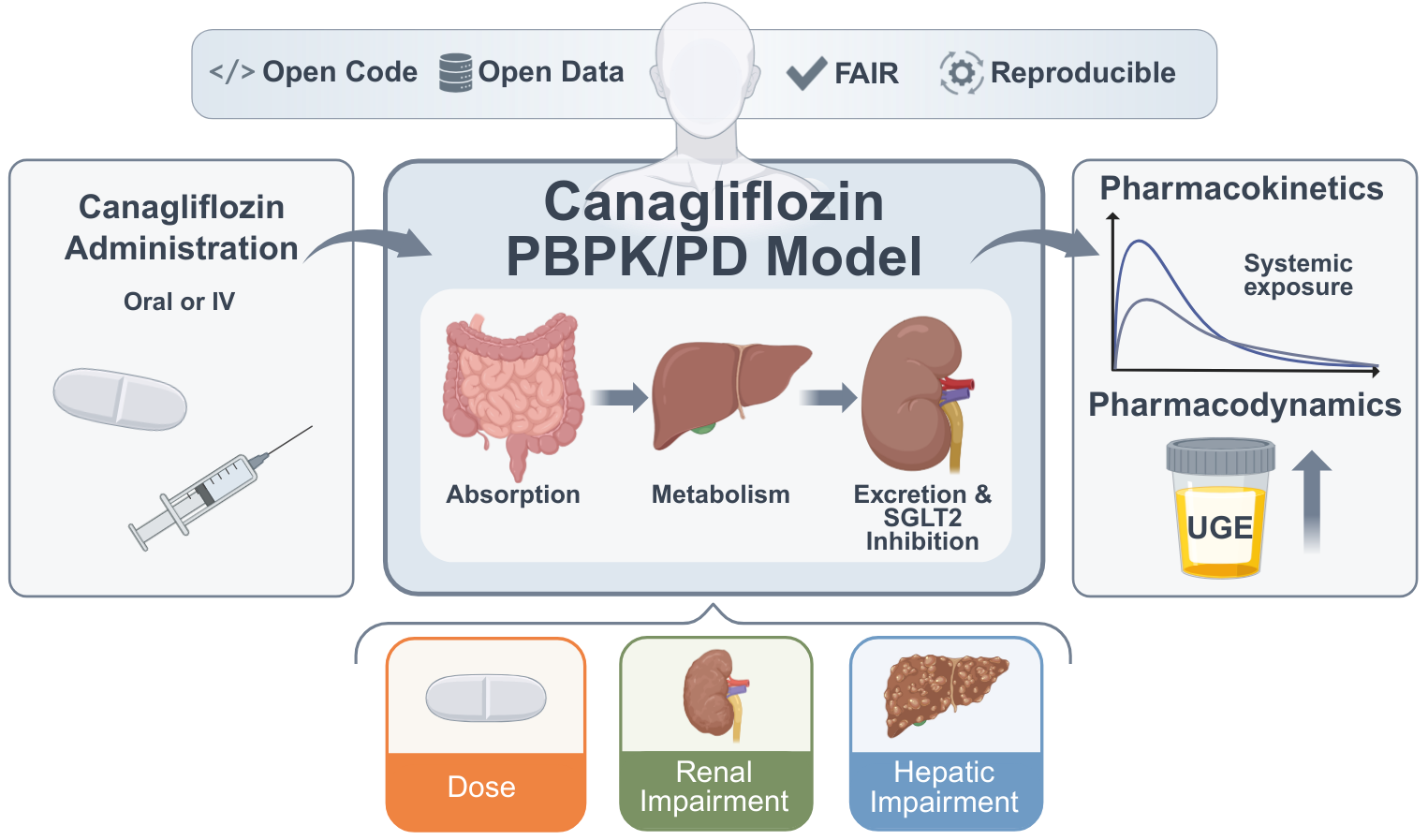 We present a whole-body PBPK/PD digital twin of canagliflozin to investigate pharmacokinetic and pharmacodynamic variability in type 2 diabetes mellitus, with a focus on renal and hepatic impairment, metabolite exposure, and dose-dependent effects.
We present a whole-body PBPK/PD digital twin of canagliflozin to investigate pharmacokinetic and pharmacodynamic variability in type 2 diabetes mellitus, with a focus on renal and hepatic impairment, metabolite exposure, and dose-dependent effects.
 In an interview with Matthias König, the Gesellschaft für Informatik explores how digital twins are becoming as complex as the human body itself and how they enable personalized prediction, simulation, and decision support in medicine.
In an interview with Matthias König, the Gesellschaft für Informatik explores how digital twins are becoming as complex as the human body itself and how they enable personalized prediction, simulation, and decision support in medicine.
 The Open Science Workshop 2025 brought together participants from across the Berlin University Alliance for a full-day, hands-on introduction to reproducible research, FAIR data, and open-source workflows.
The Open Science Workshop 2025 brought together participants from across the Berlin University Alliance for a full-day, hands-on introduction to reproducible research, FAIR data, and open-source workflows.
 Successful completion of the AI Teaching Certificate (200 AE) at Humboldt-Universität zu Berlin, covering AI foundations, ethics, and higher education didactics within the KI-Campus 2.0 and AI Skills Initiative.
Successful completion of the AI Teaching Certificate (200 AE) at Humboldt-Universität zu Berlin, covering AI foundations, ethics, and higher education didactics within the KI-Campus 2.0 and AI Skills Initiative.
 We offer Internships in 2026, academic three-month research stays at Humboldt-Universität zu Berlin. The application for the Summer Term Internships 2026 is open from 05 November to 18 December 2025.
We offer Internships in 2026, academic three-month research stays at Humboldt-Universität zu Berlin. The application for the Summer Term Internships 2026 is open from 05 November to 18 December 2025.
 We present a physiologically based PBPK/PD digital twin of dapagliflozin to analyze pharmacokinetic and pharmacodynamic variability in type 2 diabetes mellitus, with a focus on dosing, hepatorenal impairment, and food effects.
We present a physiologically based PBPK/PD digital twin of dapagliflozin to analyze pharmacokinetic and pharmacodynamic variability in type 2 diabetes mellitus, with a focus on dosing, hepatorenal impairment, and food effects.
 We introduce a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of losartan and its active metabolite E3174 as a digital twin to study variability in blood pressure regulation and support individualized antihypertensive therapy.
We introduce a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of losartan and its active metabolite E3174 as a digital twin to study variability in blood pressure regulation and support individualized antihypertensive therapy.
 We present a digital pharmacokinetic twin of glimepiride that simulates how patient-specific factors affect drug behavior, supporting individualized and stratified treatment of type 2 diabetes.
We present a digital pharmacokinetic twin of glimepiride that simulates how patient-specific factors affect drug behavior, supporting individualized and stratified treatment of type 2 diabetes.
 Our latest preprint presents results from a survey study exploring how clinicians in hepatology and hepatobiliary surgery perceive artificial intelligence (AI) and digital twins in clinical decision-making, highlighting both enthusiasm for innovation and concerns about trust, liability, and regulation.
Our latest preprint presents results from a survey study exploring how clinicians in hepatology and hepatobiliary surgery perceive artificial intelligence (AI) and digital twins in clinical decision-making, highlighting both enthusiasm for innovation and concerns about trust, liability, and regulation.
 Dr. Matthias König has successfully completed the comprehensive Leadership Certificate of the Berlin Leadership Academy, strengthening his leadership skills for guiding research teams and fostering effective collaboration in academia.
Dr. Matthias König has successfully completed the comprehensive Leadership Certificate of the Berlin Leadership Academy, strengthening his leadership skills for guiding research teams and fostering effective collaboration in academia.
 This thesis presents a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of losartan and its metabolite E3174, providing mechanistic insights into how physiological and genetic factors influence therapy and supporting individualized dosing strategies.
This thesis presents a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of losartan and its metabolite E3174, providing mechanistic insights into how physiological and genetic factors influence therapy and supporting individualized dosing strategies.
 Our latest preprint introduces a comprehensive physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of rivaroxaban, enabling the simulation of drug absorption, distribution, metabolism, excretion, and coagulation dynamics under diverse clinical conditions.
Our latest preprint introduces a comprehensive physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of rivaroxaban, enabling the simulation of drug absorption, distribution, metabolism, excretion, and coagulation dynamics under diverse clinical conditions.
 Isabella Tan, Olivia Yau and Minjun Kim presented there PBPK/PD modeling projects on GLP-1 agonists for Program Day here at Humboldt Universität-zu Berlin.
Isabella Tan, Olivia Yau and Minjun Kim presented there PBPK/PD modeling projects on GLP-1 agonists for Program Day here at Humboldt Universität-zu Berlin.
 Monika Jesionek has started a new ERASMUS project to develop a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of rapamycin, aiming to better understand interindividual variability and support the development of an immune digital twin.
Monika Jesionek has started a new ERASMUS project to develop a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of rapamycin, aiming to better understand interindividual variability and support the development of an immune digital twin.
 In this bachelor thesis, a PBPK/PD model of dapagliflozin was developed to investigate how physiological and external factors such as hepatorenal impairment and food intake influence drug behavior and therapeutic efficacy in type 2 diabetes treatment.
In this bachelor thesis, a PBPK/PD model of dapagliflozin was developed to investigate how physiological and external factors such as hepatorenal impairment and food intake influence drug behavior and therapeutic efficacy in type 2 diabetes treatment.
 Dr. Matthias König joined an interdisciplinary panel at the Berlin University Alliance Fellows Club to discuss how inclusive design influences research across fields. The event highlighted diverse approaches—from AI in healthcare to quantum computing—to advance pluralistic and equitable science.
Dr. Matthias König joined an interdisciplinary panel at the Berlin University Alliance Fellows Club to discuss how inclusive design influences research across fields. The event highlighted diverse approaches—from AI in healthcare to quantum computing—to advance pluralistic and equitable science.
 From June 3–6, we successfully conducted our redesigned course on Pharmacokinetic/Pharmacodynamic (PK/PD) Modelling. Developed as part of the Digital Health Profession Educator (DHPE) program, the course emphasized interactivity and hands-on learning in the context of clinical pharmacology.
From June 3–6, we successfully conducted our redesigned course on Pharmacokinetic/Pharmacodynamic (PK/PD) Modelling. Developed as part of the Digital Health Profession Educator (DHPE) program, the course emphasized interactivity and hands-on learning in the context of clinical pharmacology.
 Matthias König has been awarded €15,000 in startup funding from the DFG Priority Programme SPP2311 to develop SPP-FEMVis, a web-based visualization platform for Finite Element Method (FEM) simulations supporting open science and interdisciplinary collaboration.
Matthias König has been awarded €15,000 in startup funding from the DFG Priority Programme SPP2311 to develop SPP-FEMVis, a web-based visualization platform for Finite Element Method (FEM) simulations supporting open science and interdisciplinary collaboration.
 At the Festival della Robotica 2025 in Pisa, Mariia Myshkina presented our research from the Algonomy project, highlighting how AI and Digital Twins can support decision-making in hepatobiliary surgery.
At the Festival della Robotica 2025 in Pisa, Mariia Myshkina presented our research from the Algonomy project, highlighting how AI and Digital Twins can support decision-making in hepatobiliary surgery.
 We’re excited to announce new research projects in our group focusing on personalized treatment strategies for diabetes and improved anesthesia management through computational modeling and control systems.
We’re excited to announce new research projects in our group focusing on personalized treatment strategies for diabetes and improved anesthesia management through computational modeling and control systems.
 In an interview for the Berlin University Alliance, Matthias König discusses the reproducibility crisis in biology and the transformative potential of Open Science. As a BUA Open Science Ambassador, he highlights the importance of FAIR data, open standards, and cultural change in academia.
In an interview for the Berlin University Alliance, Matthias König discusses the reproducibility crisis in biology and the transformative potential of Open Science. As a BUA Open Science Ambassador, he highlights the importance of FAIR data, open standards, and cultural change in academia.
 Michelle Elias has successfully completed her Bachelor thesis on a physiologically based pharmacokinetic (PBPK) model of glimepiride, focusing on interindividual variability, hepatorenal impairment, and pharmacogenomics.
Michelle Elias has successfully completed her Bachelor thesis on a physiologically based pharmacokinetic (PBPK) model of glimepiride, focusing on interindividual variability, hepatorenal impairment, and pharmacogenomics.
 Matthias König elected as an SBML Editor for the 2025–2027 term.
Matthias König elected as an SBML Editor for the 2025–2027 term.
 As a featured BUA Fellow at Humboldt-Universität, Matthias König advances digital medicine and systems biology at the intersection of data, modeling, and clinical research.
As a featured BUA Fellow at Humboldt-Universität, Matthias König advances digital medicine and systems biology at the intersection of data, modeling, and clinical research.
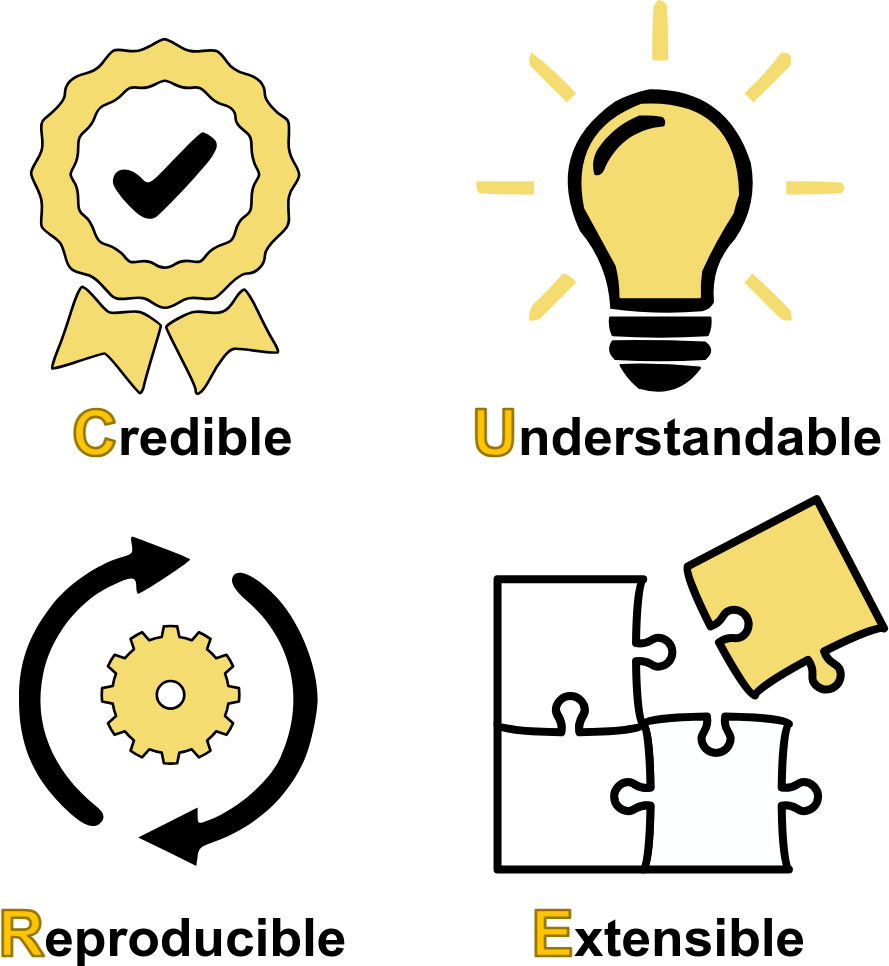 Our latest preprint, From FAIR to CURE: Guidelines for Computational Models of Biological Systems, introduces the CURE principles—Credible, Understandable, Reproducible, and Extensible—as a framework to improve the development and use of computational models in biology and medicine.
Our latest preprint, From FAIR to CURE: Guidelines for Computational Models of Biological Systems, introduces the CURE principles—Credible, Understandable, Reproducible, and Extensible—as a framework to improve the development and use of computational models in biology and medicine.
 We are excited to announce that Mariia Myshkina, Elisabetta Casabianca, and Matthias König participated in the EP PerMed Conference on Personalised Medicine Research, taking place on February 11–12, 2025, in Berlin, Germany.
We are excited to announce that Mariia Myshkina, Elisabetta Casabianca, and Matthias König participated in the EP PerMed Conference on Personalised Medicine Research, taking place on February 11–12, 2025, in Berlin, Germany.
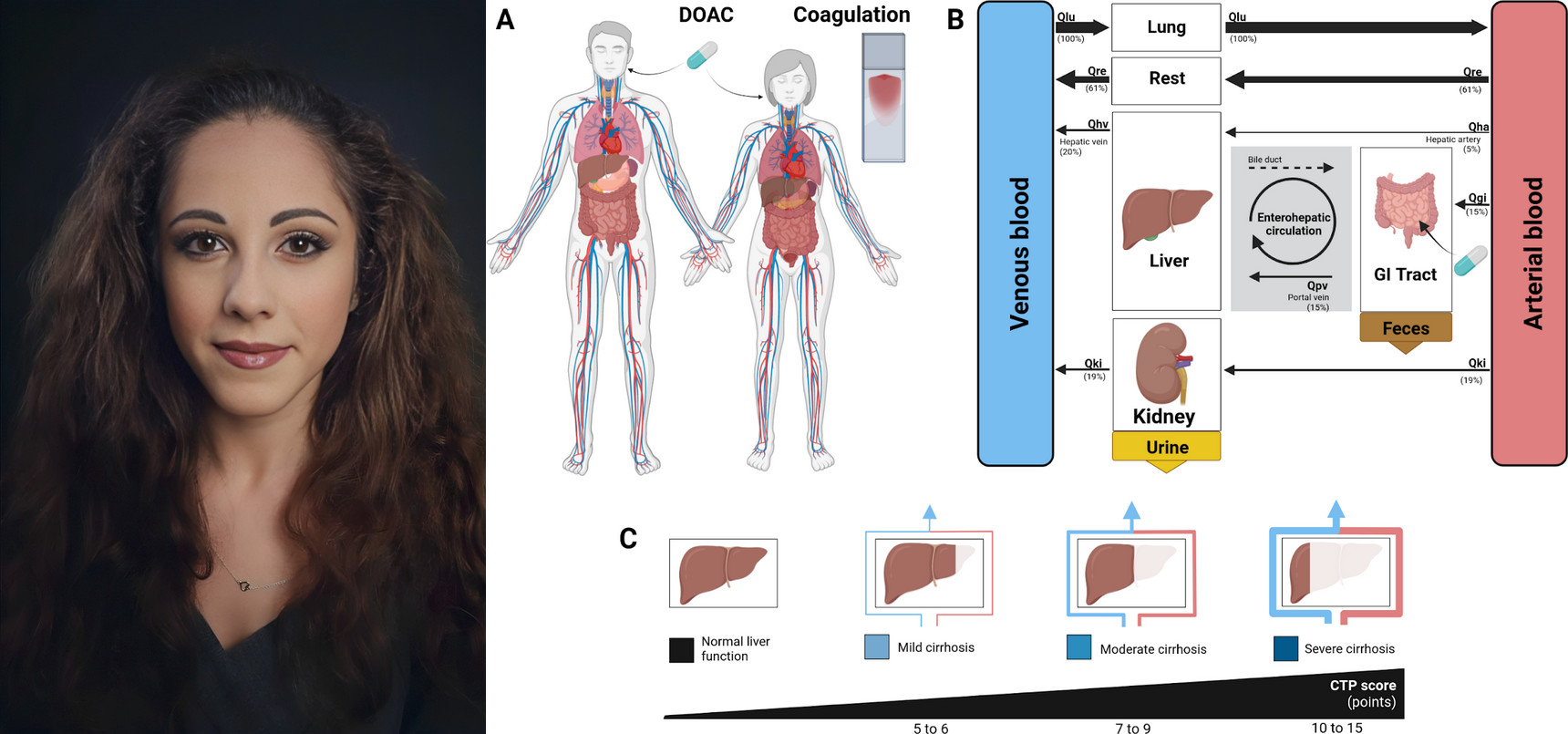 PBPK Model Development for Tirzepatide in Diabetes and Obesity Treatment
PBPK Model Development for Tirzepatide in Diabetes and Obesity Treatment
 PBPK Model Development for Tirzepatide in Diabetes and Obesity Treatment
PBPK Model Development for Tirzepatide in Diabetes and Obesity Treatment
 Circle U. has funded AlgoNomy: Advancing Doctor-Patient Autonomy in AI-Driven Healthcare, a collaborative project tackling the issue of digital paternalism—where algorithms dominate healthcare decisions over human collaboration. Partnering with six leading European universities, AlgoNomy aims to develop ethical, transparent, and patient-centered AI systems that preserve autonomy and promote shared decision-making.
Circle U. has funded AlgoNomy: Advancing Doctor-Patient Autonomy in AI-Driven Healthcare, a collaborative project tackling the issue of digital paternalism—where algorithms dominate healthcare decisions over human collaboration. Partnering with six leading European universities, AlgoNomy aims to develop ethical, transparent, and patient-centered AI systems that preserve autonomy and promote shared decision-making.
 We offer Internships in 2025, academic three-month research stays at Humboldt-Universität zu Berlin. The application for the Summer Term Internships 2025 is open from 11 November to 20 December 2024.
We offer Internships in 2025, academic three-month research stays at Humboldt-Universität zu Berlin. The application for the Summer Term Internships 2025 is open from 11 November to 20 December 2024.
 Ennie Tensil and Michelle Elias start Bachelor projects on losartan and glimeperide
Ennie Tensil and Michelle Elias start Bachelor projects on losartan and glimeperide
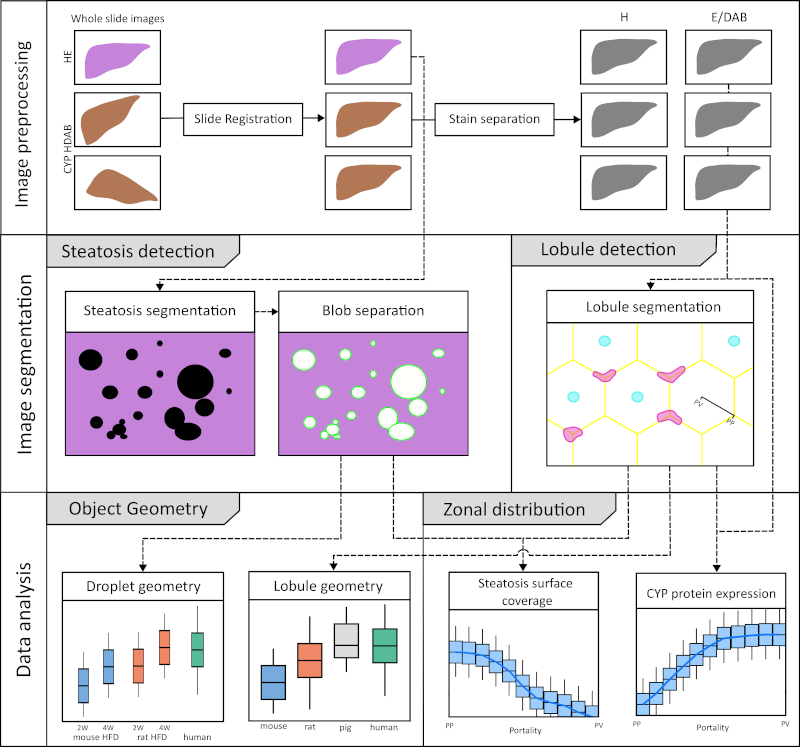 Jonas Küttner submitted his Master thesis developing a workflow for image analysis in the liver
Jonas Küttner submitted his Master thesis developing a workflow for image analysis in the liver
 Proud to be selected as Open Science Ambassador of the Berlin University Alliance (BUA) in 2024.
Proud to be selected as Open Science Ambassador of the Berlin University Alliance (BUA) in 2024.
 Matthias König selected for the Digital Health Professions Educator (DHPE) Program
Matthias König selected for the Digital Health Professions Educator (DHPE) Program
 Proud to participate in Humboldt Internship Program 2024. Great work by Vera, Deepa and Prerna.
Proud to participate in Humboldt Internship Program 2024. Great work by Vera, Deepa and Prerna.
 Proud to participate in the final ecosystem meeting of EDITH on building the virtual human twin (VHT) in Amsterdam.
Proud to participate in the final ecosystem meeting of EDITH on building the virtual human twin (VHT) in Amsterdam.
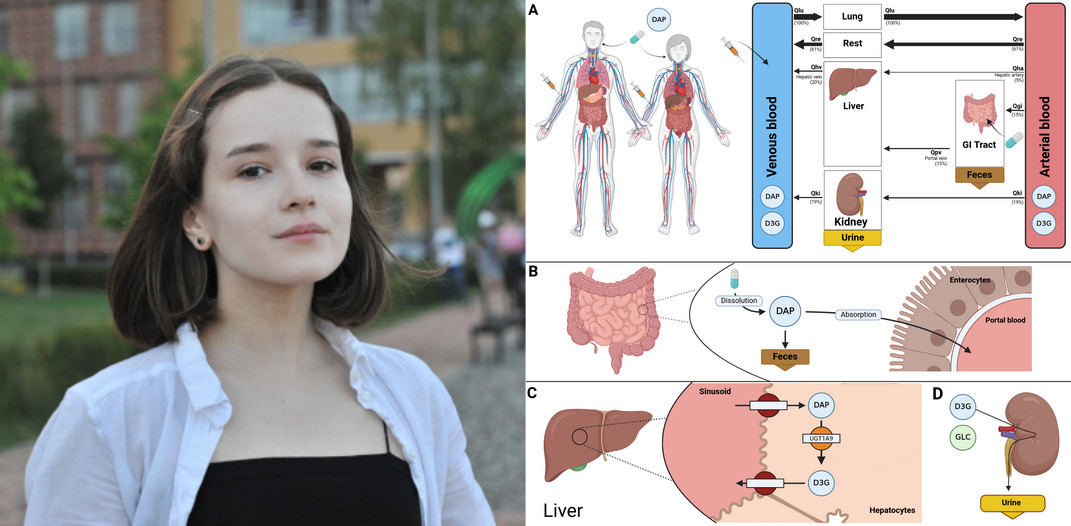 Vera Tereshchuk will develop a PBPK models for canagliflozin, aiming to enhance understanding of pharmacokinetics and pharmacodynamics in diabetes.
Vera Tereshchuk will develop a PBPK models for canagliflozin, aiming to enhance understanding of pharmacokinetics and pharmacodynamics in diabetes.
 Mariia Myshkina and Matthias König presented our captopril modeling results during the PAGE2024 (Population Approach Group Europe) meeting in Rome.
Mariia Myshkina and Matthias König presented our captopril modeling results during the PAGE2024 (Population Approach Group Europe) meeting in Rome.
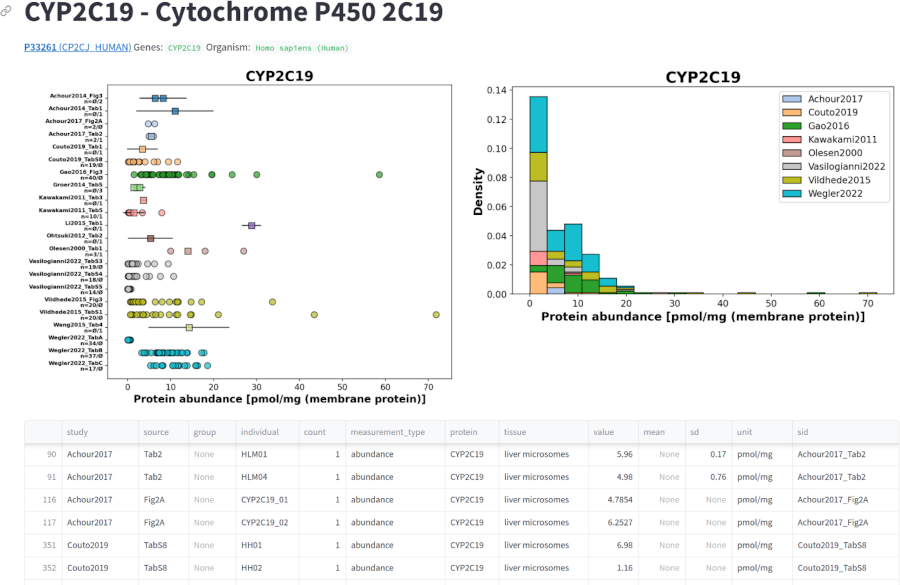 Afruja Hossain finished her Bachelor thesis developing a database of CYP and UGT enzymes
Afruja Hossain finished her Bachelor thesis developing a database of CYP and UGT enzymes
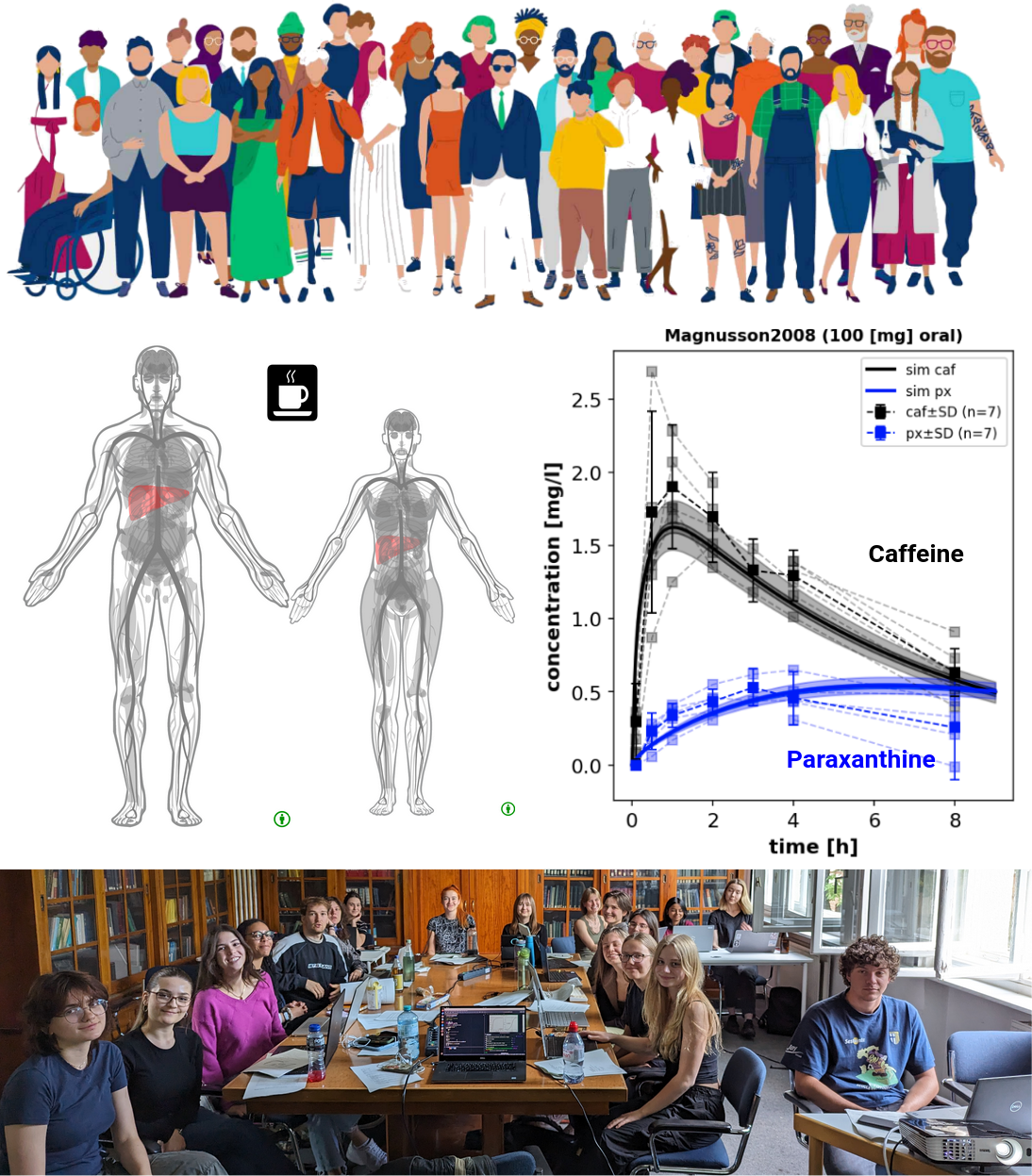 We are excited to announce the successful completion of our one-week intensive course on Pharmacokinetic Modelling, where students delved into principles, drug distribution, clearance, elimination, and various modelling techniques to enhance drug therapy and development.
We are excited to announce the successful completion of our one-week intensive course on Pharmacokinetic Modelling, where students delved into principles, drug distribution, clearance, elimination, and various modelling techniques to enhance drug therapy and development.
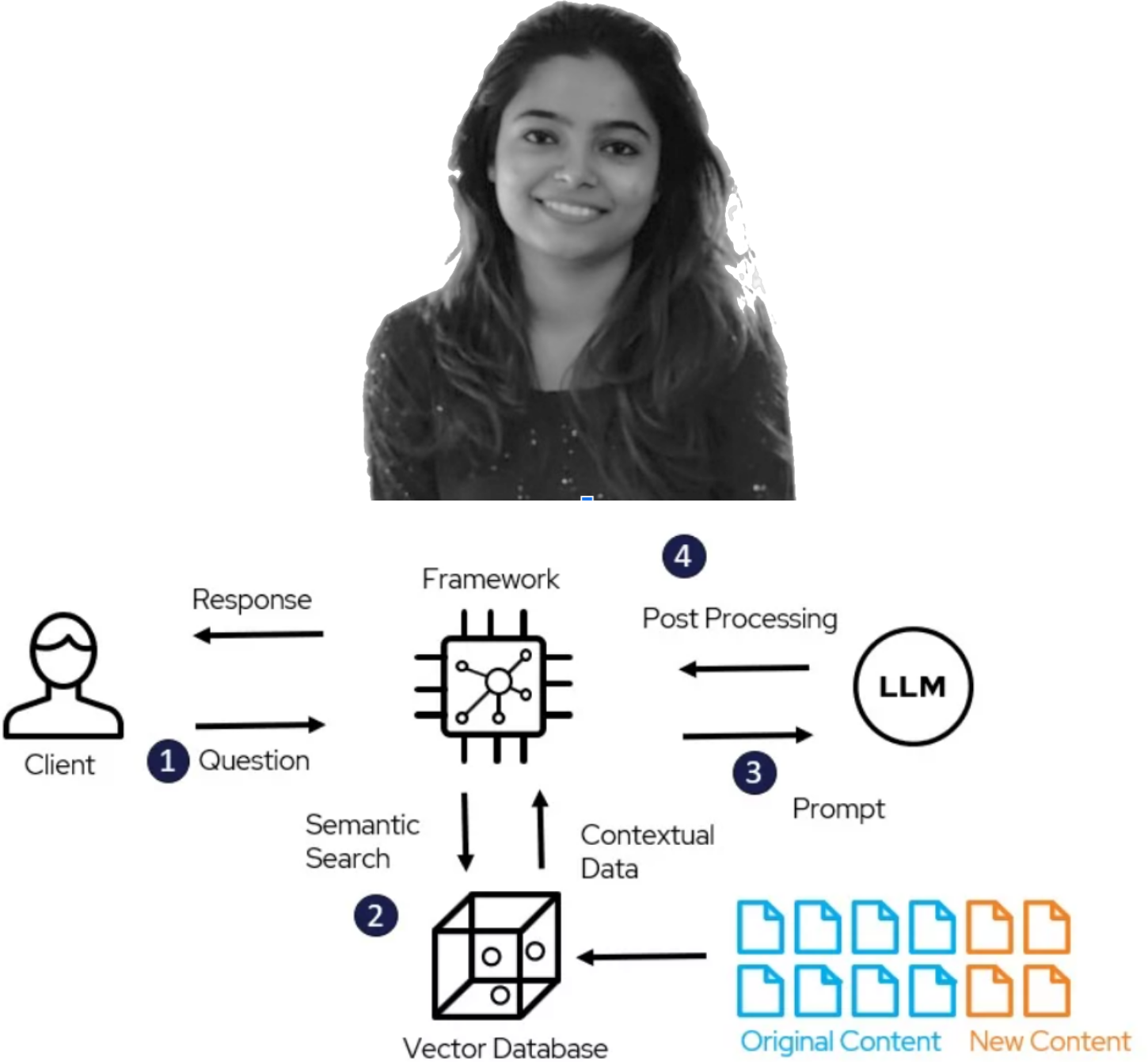 We are excited to announce the start of a new project under the Humboldt Internship Program (HIP) by Prerna Parakkat. Prerna will be working on PK-LLM: Implementing a Large Language Model (LLM) as an Expert System for Pharmacokinetic (PK) Data Curation from Scientific Publications.
We are excited to announce the start of a new project under the Humboldt Internship Program (HIP) by Prerna Parakkat. Prerna will be working on PK-LLM: Implementing a Large Language Model (LLM) as an Expert System for Pharmacokinetic (PK) Data Curation from Scientific Publications.
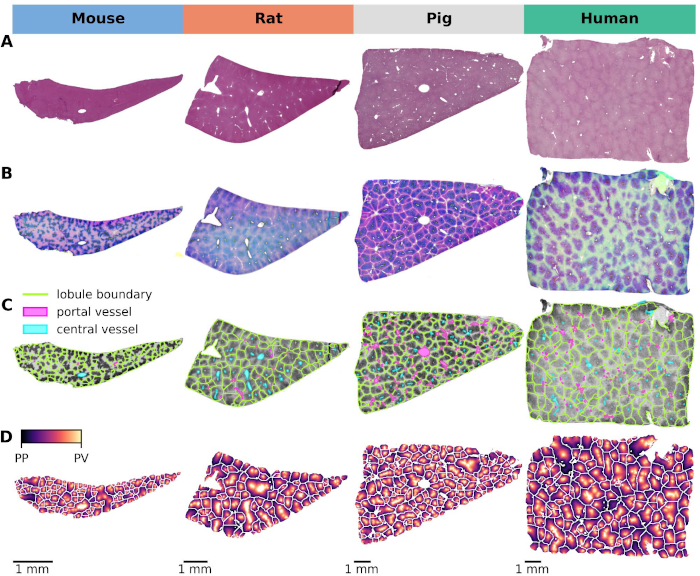 Our latest publication on 'Cross-Species Variability in Lobular Geometry & Cytochrome P450 Hepatic Zonation' reveals insights into liver function across species using WSI technology.
Our latest publication on 'Cross-Species Variability in Lobular Geometry & Cytochrome P450 Hepatic Zonation' reveals insights into liver function across species using WSI technology.
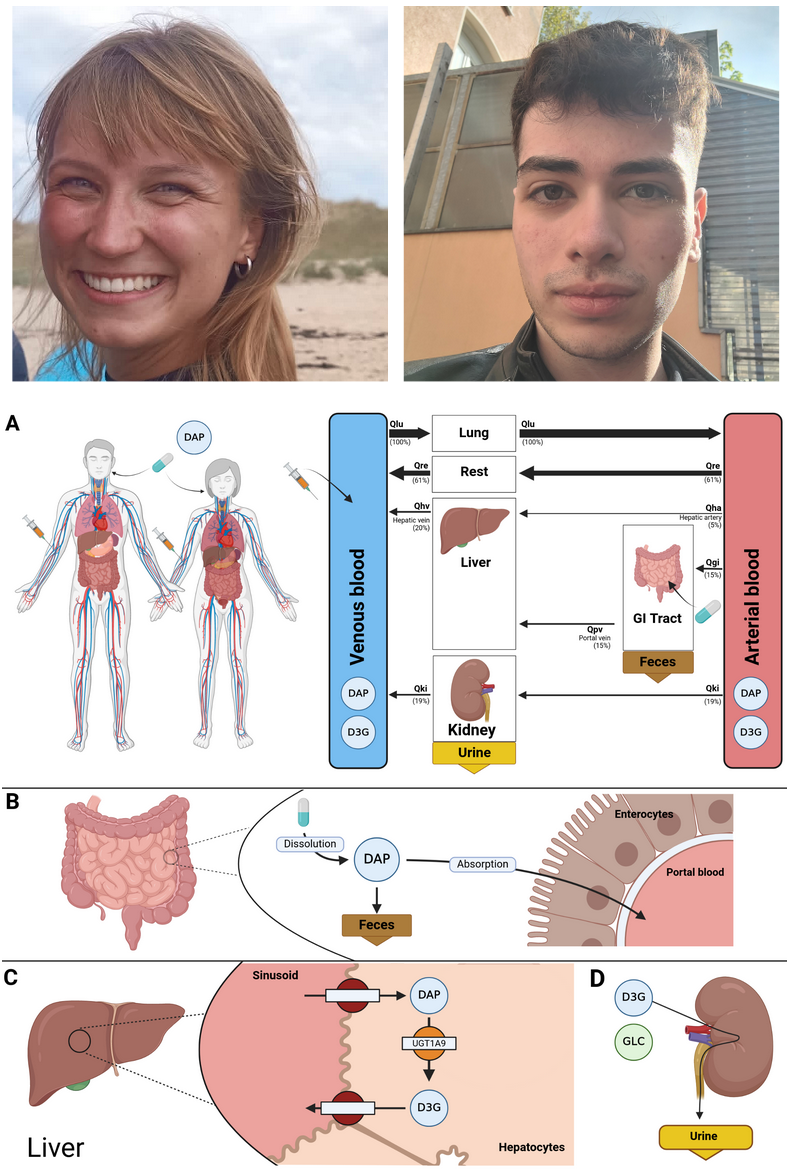 Nike Nemitz and Yusuf Ali Kulanoglu will develop PBPK models for dapagliflozin and aliskiren, respectively, aiming to enhance understanding of pharmacokinetics and pharmacodynamics in diabetes and cardiovascular treatments.
Nike Nemitz and Yusuf Ali Kulanoglu will develop PBPK models for dapagliflozin and aliskiren, respectively, aiming to enhance understanding of pharmacokinetics and pharmacodynamics in diabetes and cardiovascular treatments.
 Level 1 Version 5 of SED-ML enhances the ability to annotate, archive, share, and reproduce computational simulation experiments by allowing modelers to define tasks, model changes, ranges, and outputs using the Kinetic Simulation Algorithm Ontology (KiSAO).
Level 1 Version 5 of SED-ML enhances the ability to annotate, archive, share, and reproduce computational simulation experiments by allowing modelers to define tasks, model changes, ranges, and outputs using the Kinetic Simulation Algorithm Ontology (KiSAO).
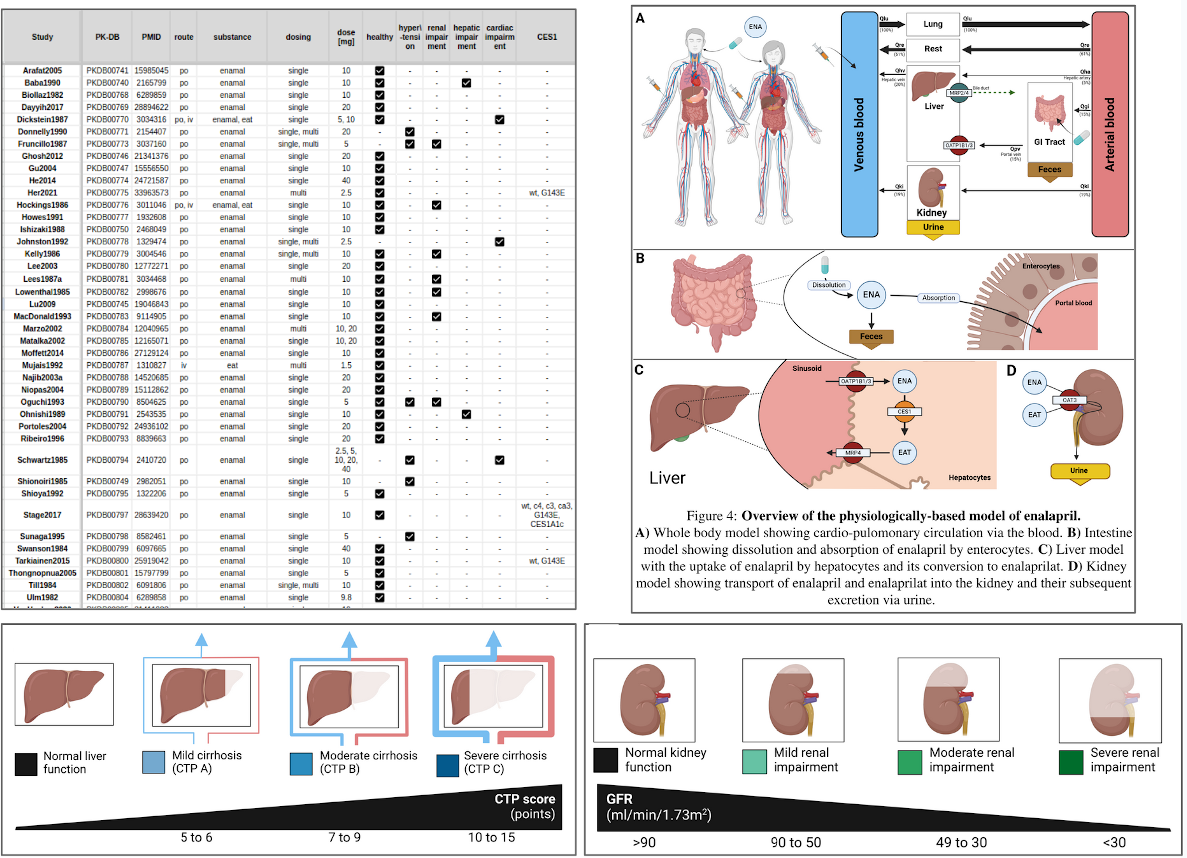 Shubhankar Palwankar defended his Master Thesis developing a PBPK model of enalapril
Shubhankar Palwankar defended his Master Thesis developing a PBPK model of enalapril
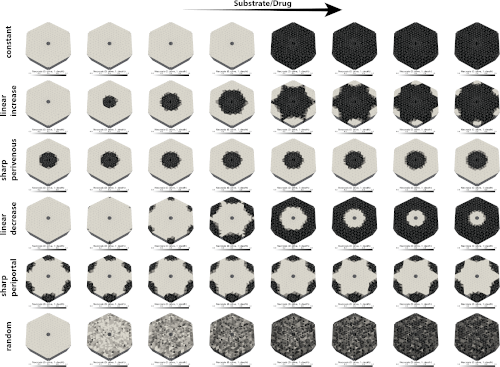 We're excited to announce the preprint of our latest research, 'Simulation of zonation-function relationships in the liver using coupled multiscale models: Application to drug-induced liver injury.' This study introduces a multiscale digital twin of the liver, integrating models from cellular to lobular scales, and sheds light on how liver zonation patterns influence drug-induced liver injury.
We're excited to announce the preprint of our latest research, 'Simulation of zonation-function relationships in the liver using coupled multiscale models: Application to drug-induced liver injury.' This study introduces a multiscale digital twin of the liver, integrating models from cellular to lobular scales, and sheds light on how liver zonation patterns influence drug-induced liver injury.
 New Internship Project Launch: Development of a Physiologically Based Pharmacokinetics Model of Morphine. Deepa supported by Rohini and Rohit will develop a PBPK model of morphine.
New Internship Project Launch: Development of a Physiologically Based Pharmacokinetics Model of Morphine. Deepa supported by Rohini and Rohit will develop a PBPK model of morphine.
 We welcome Mariia to our team who will be working on computational modeling of the liver within ATLAS.
We welcome Mariia to our team who will be working on computational modeling of the liver within ATLAS.
 We participated in the first EDITH-CSA 'Ecosystem meeting on building the Virtual Human Twin was held at the École Polytechnique de Paris on January 18 and 19, 2024.
We participated in the first EDITH-CSA 'Ecosystem meeting on building the Virtual Human Twin was held at the École Polytechnique de Paris on January 18 and 19, 2024.
 We are pleased to announce our latest paper in eLife that provides a roadmap for integrating reproducible research and open science practices into academic work.
We are pleased to announce our latest paper in eLife that provides a roadmap for integrating reproducible research and open science practices into academic work.
 Advancing Precision Nutrition: Workshop Success on Dietary Digital Twin, a workshop in Leiden from 30 October - 3 November 2023.
Advancing Precision Nutrition: Workshop Success on Dietary Digital Twin, a workshop in Leiden from 30 October - 3 November 2023.
 We successfully presented our research on reproducible digital twins for personalized liver function assessment at the e:Med Meeting 2023 on Systems Medicine.
We successfully presented our research on reproducible digital twins for personalized liver function assessment at the e:Med Meeting 2023 on Systems Medicine.
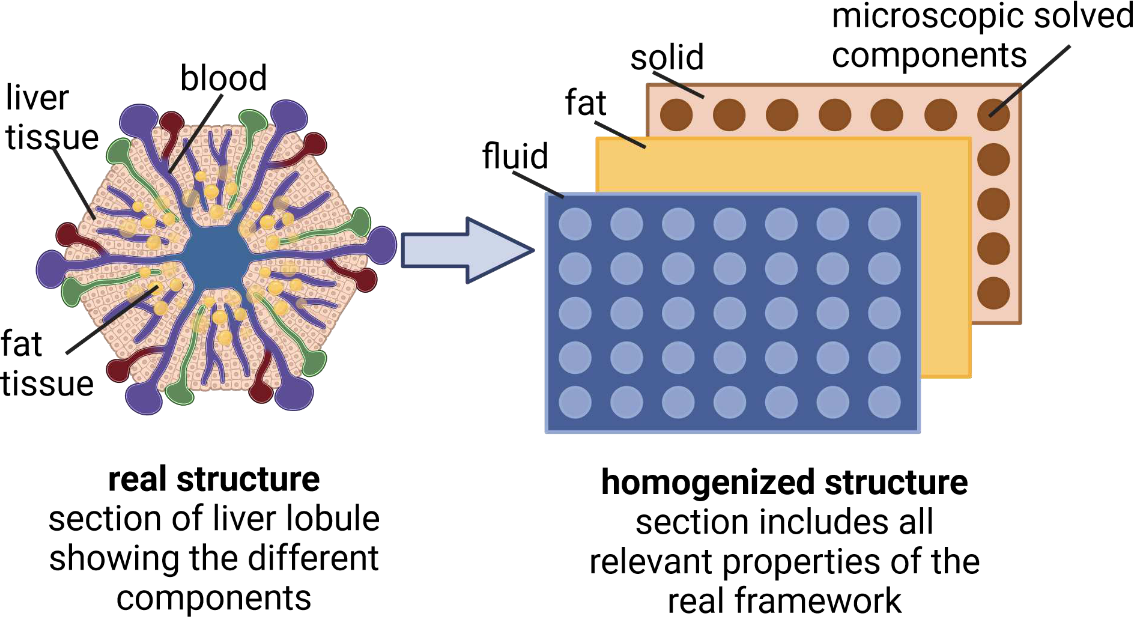 Our latest preprint is now out: An integrated multiscale model for quantifying fat zonation in liver lobules.
Our latest preprint is now out: An integrated multiscale model for quantifying fat zonation in liver lobules.
 We welcome Hannah Menghis who will establish a PBPK model of semaglutide.
We welcome Hannah Menghis who will establish a PBPK model of semaglutide.
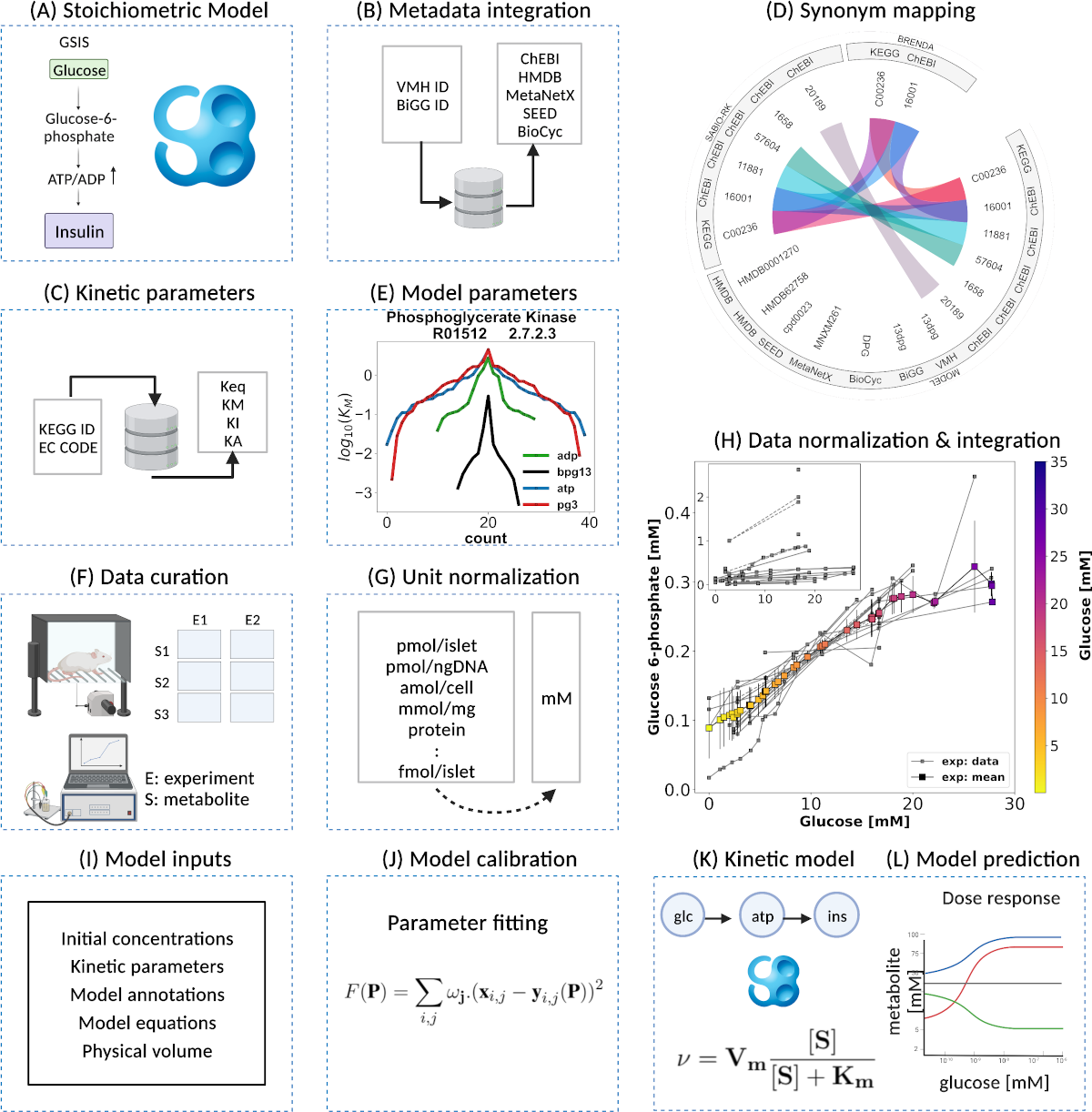 Our latest publication is now on Frontiers of Endocrinology: A pathway model of glucose-stimulated insulin secretion in the pancreatic β-cell
Our latest publication is now on Frontiers of Endocrinology: A pathway model of glucose-stimulated insulin secretion in the pancreatic β-cell
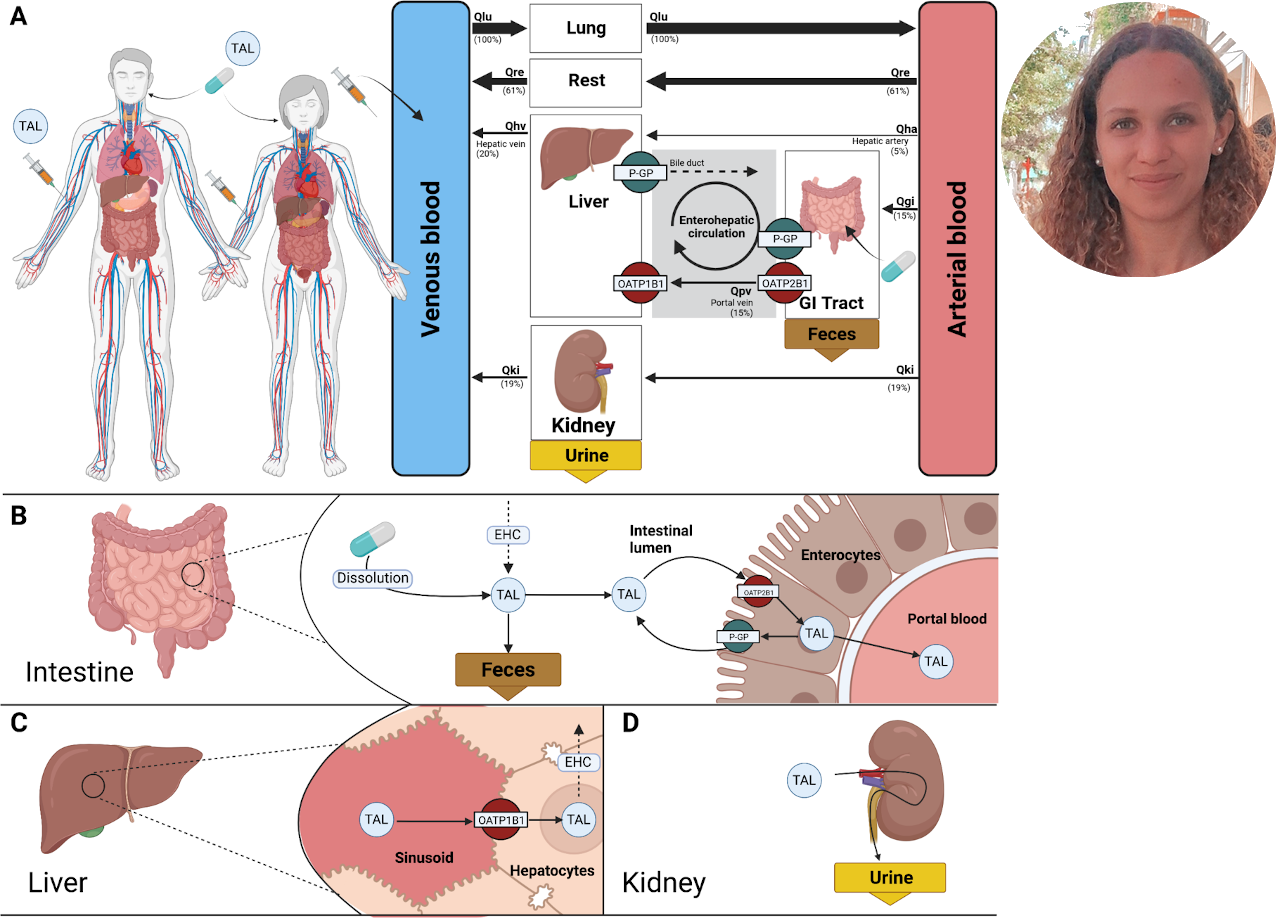 Beatrice Stemmer Mallol submitted her Bachelor thesis developing a PBPK model of talinolol
Beatrice Stemmer Mallol submitted her Bachelor thesis developing a PBPK model of talinolol
 Jan Grzegorzewski defended his PhD thesis on physiologically based pharmacokinetic modeling (PBPK) for dynamical liver function tests and Cytochrome P450 (CYP) phenotyping.
Jan Grzegorzewski defended his PhD thesis on physiologically based pharmacokinetic modeling (PBPK) for dynamical liver function tests and Cytochrome P450 (CYP) phenotyping.
 Humboldt Internship Program 2023 - Computational Modeling of Drug Detoxification - A Systems Medicine Approach
Humboldt Internship Program 2023 - Computational Modeling of Drug Detoxification - A Systems Medicine Approach
 Keynote talk at the Workshop on Computational Models in Biology and Medicine 2023 organized by Nicole Radde and Sebastian Höpfl.
Keynote talk at the Workshop on Computational Models in Biology and Medicine 2023 organized by Nicole Radde and Sebastian Höpfl.
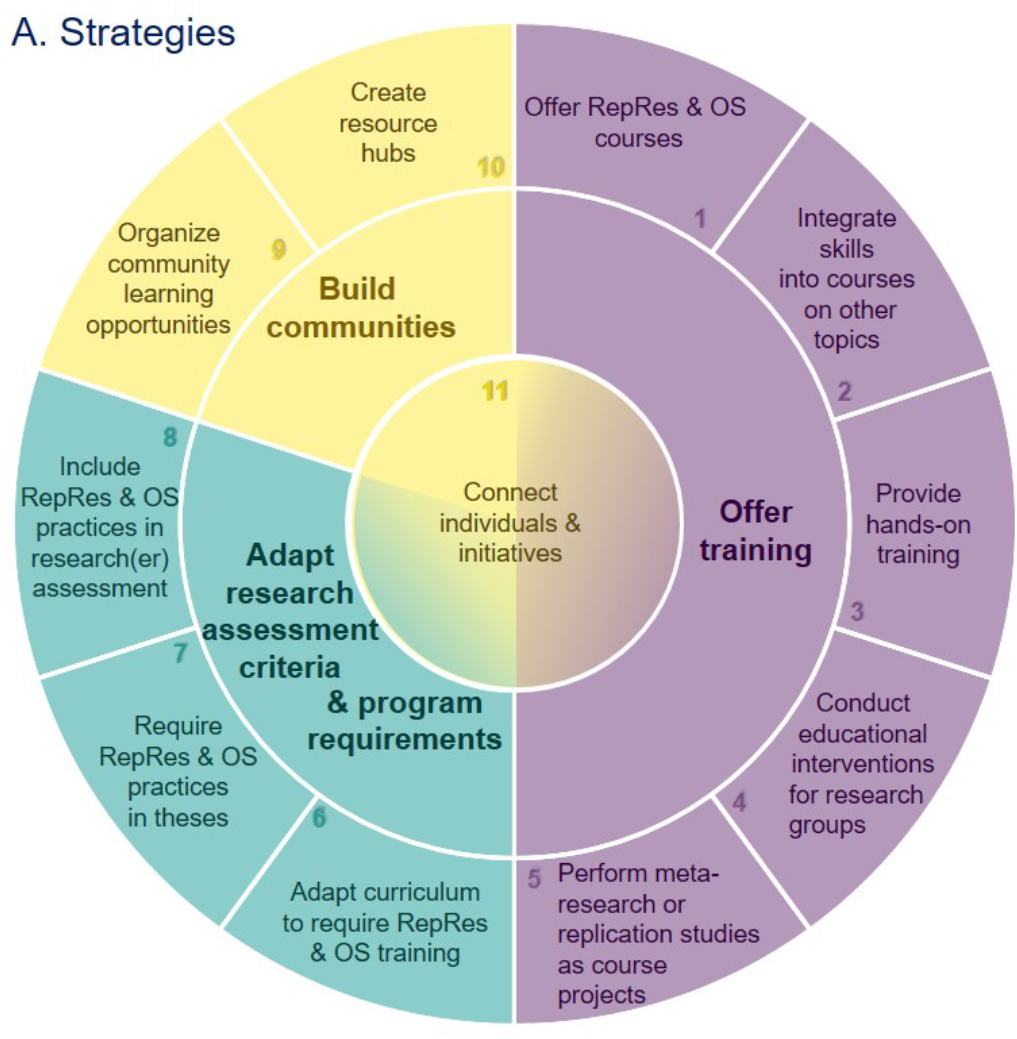 We present 11 strategies for making #OpenScience & #ReproducibleResearch the norm at research institutions, with tips for implementation & resources. Let's join hands to make these practices standard in the scientific community!
We present 11 strategies for making #OpenScience & #ReproducibleResearch the norm at research institutions, with tips for implementation & resources. Let's join hands to make these practices standard in the scientific community!
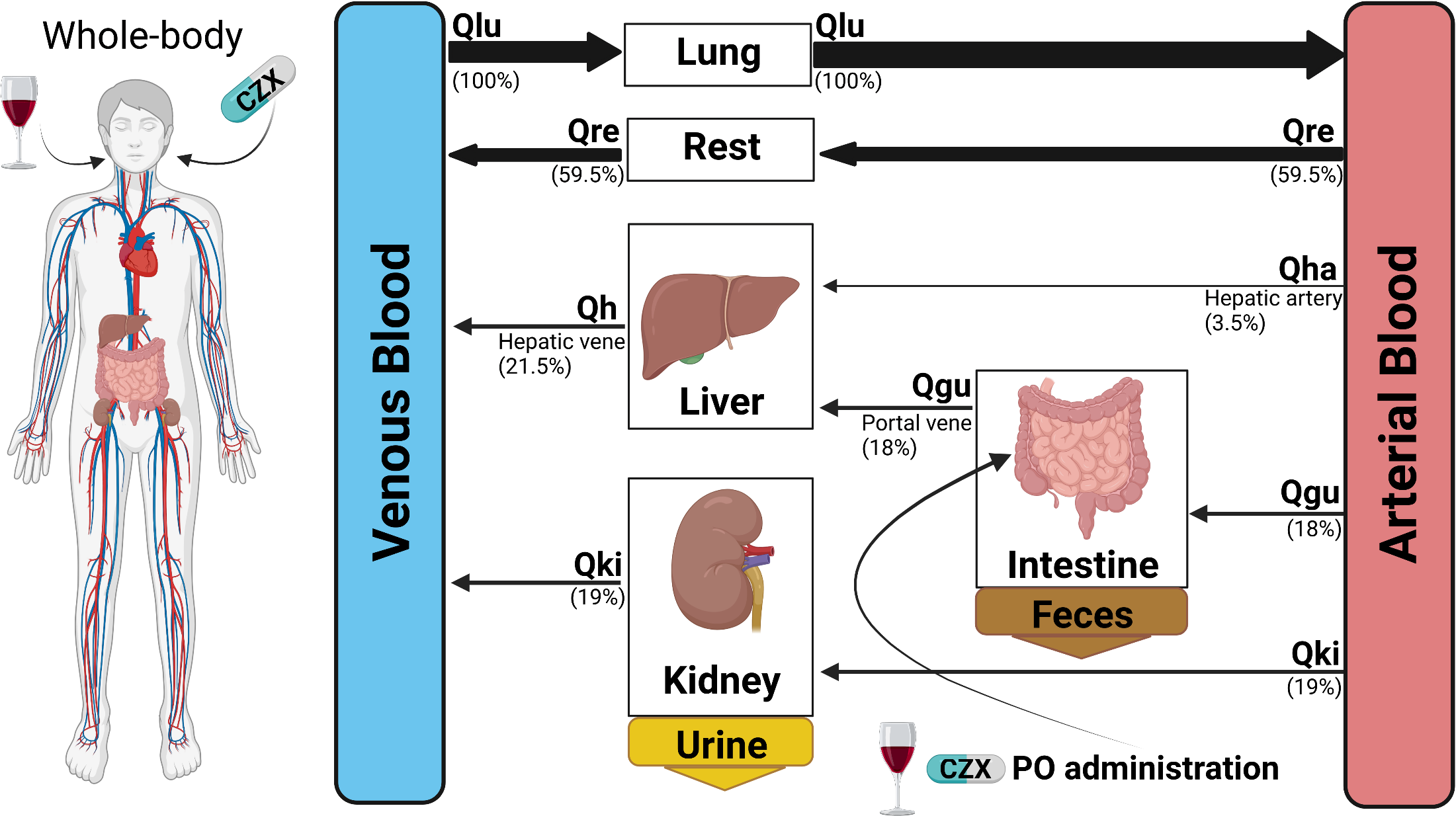 Our latest preprint is now on bioRxiv: A physiologically based pharmacokinetic model for CYP2E1 phenotyping via chlorzoxazone
Our latest preprint is now on bioRxiv: A physiologically based pharmacokinetic model for CYP2E1 phenotyping via chlorzoxazone
 M. Albadry, S. Höpfl, and M. König won the QuaLiPerF Michael Stifel Price for interdisciplinary, data-driven research.
M. Albadry, S. Höpfl, and M. König won the QuaLiPerF Michael Stifel Price for interdisciplinary, data-driven research.
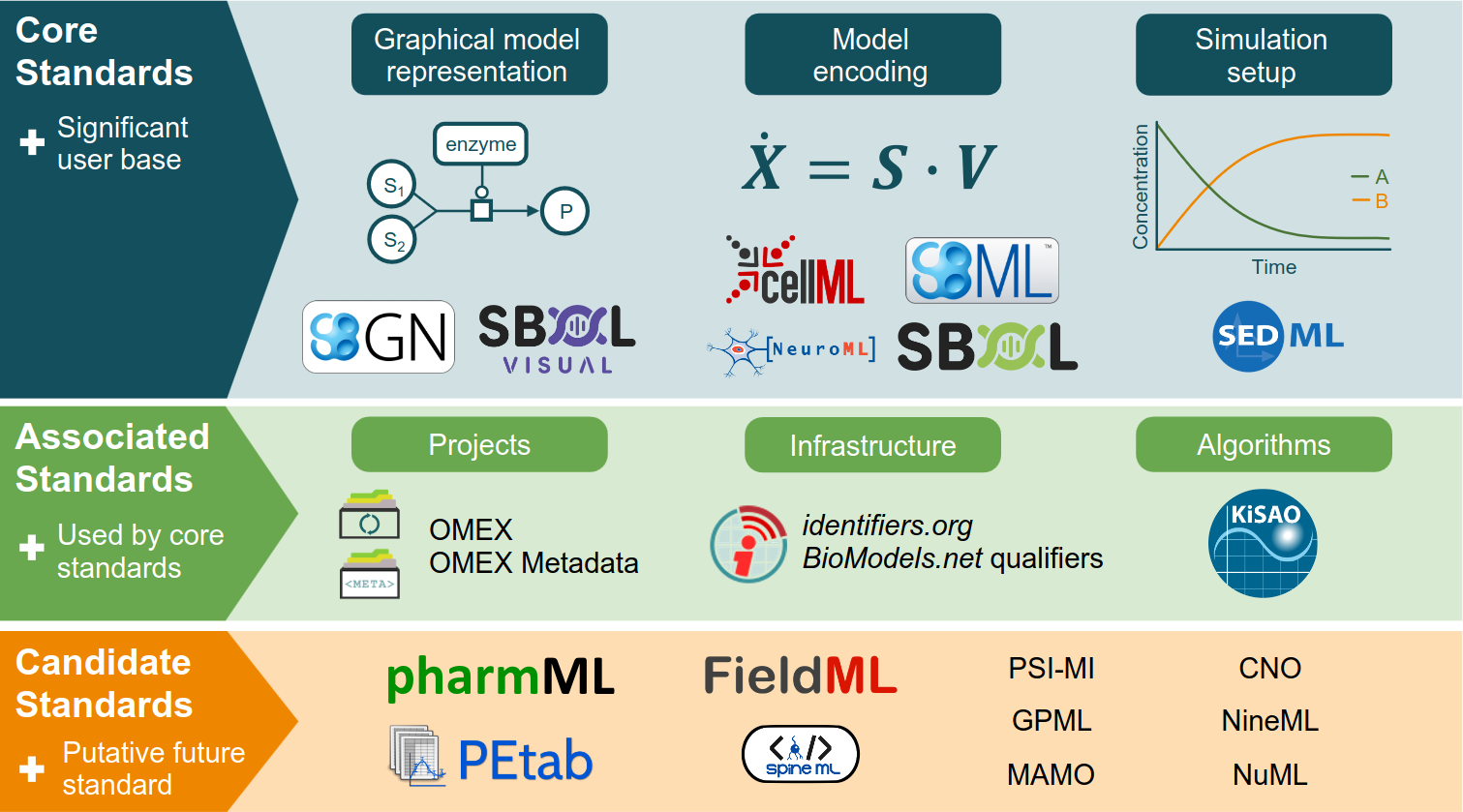 Our overview of specifications in systems and synthetic biology with a brief overview of the COMBINE 2022 is available.
Our overview of specifications in systems and synthetic biology with a brief overview of the COMBINE 2022 is available.
 Frances Okibedi and Narudeem Sojimade will develop models of the anticancer drug sorafenib in HCC and the liver contrast agent Gd-EOB-DTPA.
Frances Okibedi and Narudeem Sojimade will develop models of the anticancer drug sorafenib in HCC and the liver contrast agent Gd-EOB-DTPA.
 M. König receives funding of the Berlin University Alliance and the DFG for the X-Student Research Group: Digital twins for the treatment of hypertension
M. König receives funding of the Berlin University Alliance and the DFG for the X-Student Research Group: Digital twins for the treatment of hypertension
 Proud to be elected editor for PEtab, a data format for specifying parameter estimation problems in systems biology.
Proud to be elected editor for PEtab, a data format for specifying parameter estimation problems in systems biology.
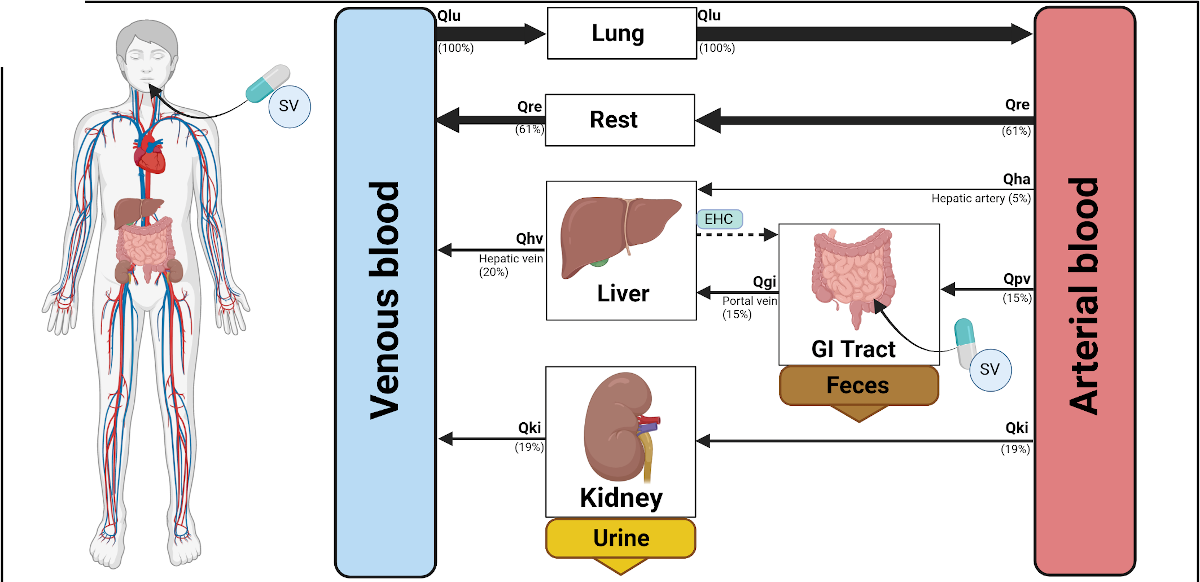 Our latest preprint is now on medRxiv: Simvastatin therapy in different subtypes of hypercholesterolemia - a physiologically based modelling approach.
Our latest preprint is now on medRxiv: Simvastatin therapy in different subtypes of hypercholesterolemia - a physiologically based modelling approach.
 The BMBF is funding our joint project ATLAS in the "Computational Life Sciences - AI Methods for Systems Medicine".
The BMBF is funding our joint project ATLAS in the "Computational Life Sciences - AI Methods for Systems Medicine".
 Afruja Hossain started working in our lab on a systematic overview of the protein variability of cytochrome P450 (CYP450) isoforms in the human liver.
Afruja Hossain started working in our lab on a systematic overview of the protein variability of cytochrome P450 (CYP450) isoforms in the human liver.
 We were selected to participate in an European policy journey on the topic EU research funding and science communication by the Berlin University Alliance (BUA).
We were selected to participate in an European policy journey on the topic EU research funding and science communication by the Berlin University Alliance (BUA).