We are actively involved in the standardization and reproducibility efforts in Systems Biology and Systems Medicine.
|


|
 |
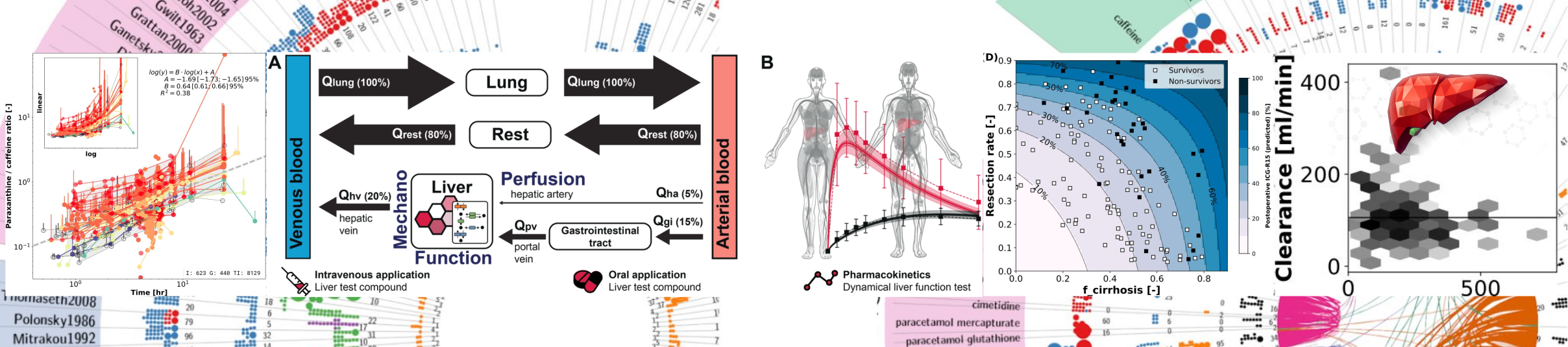
PK-DB - Pharmacokinetics database
Developed the first FAIR-compliant open database for pharmacokinetics, integrating clinical and pre-clinical trial data. PK-DB enables reproducible PBPK/PD modeling, supports individualized simulations, and has become a key infrastructure for computational pharmacology research. |
|


|
 |
sbmlutils - Python utilities for SBML
Created a versatile Python library to streamline the use of SBML models, providing robust utilities for model handling, analysis, and integration with libSBML. Widely used in reproducible modeling workflows across systems biology. |
|


|
 |
SBML4Humans - SBML simulation made easy
Designed an interactive reporting framework that makes SBML models human-readable and accessible, enabling experts and newcomers to explore model content without technical barriers. |
|


|
 |
sbmlsim - SBML simulation made easy
Built a lightweight Python package that simplifies simulations of SBML models on top of libRoadRunner, lowering the entry barrier for model testing and teaching. |
|


|
 |
cysbml - Cytoscape 3 app for the Systems Biology Markup Language
Developed and maintained a widely used Cytoscape app for visualization of SBML models in network contexts. cy3sbml has facilitated intuitive exploration of complex models in systems biology and bioinformatics. |
|


|
 |
brendapy - BRENDA in python
Authored a Python package providing direct programmatic access to BRENDA enzyme information, enabling large-scale, automated enzyme analysis and integration into computational pipelines. |
|


|
 |
libsbgnpy - Python library for SBGN
Developed a Python library for working with Systems Biology Graphical Notation (SBGN), supporting standardized visualization and integration of pathway information. |
|


|
 |
roadrunner - High-performance simulator for SBML
Contributed to the development of libRoadRunner, a C/C++ library using LLVM for ultra-fast simulation of SBML models, setting a benchmark for performance in computational biology. |
|


|
 |
COBRApy - COBRA python package
Advanced COBRApy, the leading Python package for constraint-based reconstruction and analysis, widely adopted in genome-scale metabolic modeling. Provides access to key methods such as flux balance and flux variability analysis. |
|


|
 |
tellurium - systems biology simulation library
Co-developed Tellurium, a Python-based environment for reproducible dynamical modeling of biological networks. Integrated standard formats with powerful simulation libraries, enabling accessible and transparent modeling. |