| 2025-20-21 |
|
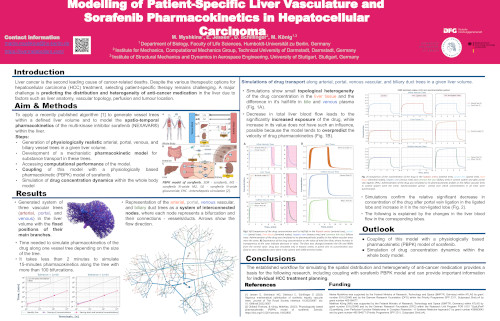
Modelling of Patient-Specific Liver Vasculature and Sorafenib Pharmacokinetics in Hepatocellular Carcinoma
Mariia Myshkina1, Etienne Jessen2, Dominik Schillinger2, Matthias König1,3
1 Department of Biology, Faculty of Life Sciences, Humboldt-Universität zu Berlin, Germany; 2 Institute for Mechanics, Computational Mechanics Group, Technical University of Darmstadt, Darmstadt, Germany; 3 Institute of Structural Mechanics and Dynamics in Aerospace Engineering, University of Stuttgart, Stuttgart, Germany
EASL Liver Cancer Summit 2026
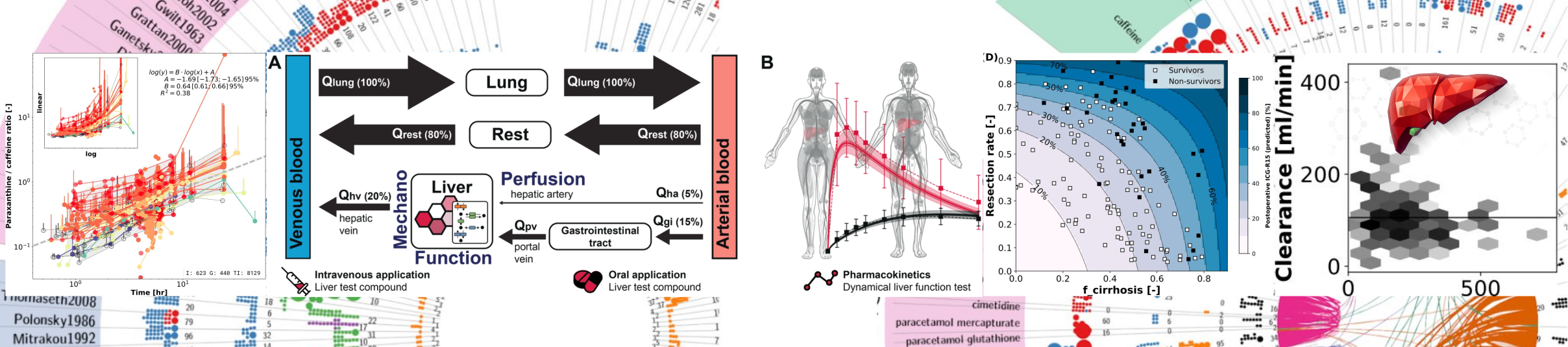
Background and Aims: Liver cancer is the second leading cause of cancer-related deaths. Despite the various therapeutic options for hepatocellular carcinoma (HCC) treatment, selecting patient-specific therapy remains challenging. A major challenge is predicting the distribution and heterogeneity of anti-cancer medication in the liver due to factors such as liver anatomy, vascular topology, perfusion and tumour location. In this study, we applied a recently published algorithm [1] to generate vessel trees within a defined liver volume and modelled the spatio-temporal pharmacokinetics of the multi-kinase inhibitor sorafenib (NEXAVAR®) within the liver. Our objectives were: (i) to generate physiologically realistic arterial, portal, venous, and biliary vessel trees in a given liver volume; (ii) to develop a mechanistic pharmacokinetic model for substance transport in these trees; (iii) to couple this model with a physiologically based pharmacokinetic (PBPK) model of sorafenib; and (iv) to simulate drug concentration dynamics within the liver. Methods: Substance transport along the vessel and biliary trees was modelled using ordinary differential equations. Each terminal segment of the trees was associated with a terminal liver volume, representing the hepatic tissue supplied by individual terminal arterial, portal, venous, and biliary segments. In each terminal volume, the local sorafenib metabolism and biliary excretion were modelled. Model performance was evaluated using published pharmacokinetic data. Results: The algorithm successfully generated physiologically realistic, non-intersecting arterial, portal, venous, and biliary trees within the liver volume. The surrogate mechanistic–PBPK model reproduced published pharmacokinetic data for sorafenib and enabled spatially resolved predictions of drug concentrations along the vascular and biliary trees and within liver tissue. Simulations revealed that the time to reach maximum sorafenib concentration in liver tissue was directly related to proximity to the start of inflow trees and inversely to proximity to the end of outflow trees. These results underscore the influence of the liver anatomy, its vascularisation topology and target location on intrahepatic drug exposure, suggesting that individualised dosing strategies could enhance therapeutic response in HCC. Conclusion: The established workflow for simulating the spatial distribution and heterogeneity of anti-cancer medication in a given liver anatomy can provide important information for individual HCC treatment planning. References [1] Jessen E, Steinbach MC, Debbaut C, Schillinger D (2022). Rigorous mathematical optimization of synthetic hepatic vascular trees. Journal of The Royal Society Interface 19:20220087. doi: 10.1098/rsif.2022.0087 [2] Okibedi Frances, & König, Matthias. (2023). Physiologically based pharmacokinetic (PBPK) model of sorafenib. Zenodo. https://doi.org/10.5281/zenodo.10432855
Keywords:
pharmacokinetics,
sorafenib,
vessel trees,
|
|
| 2025-07-07 |
|
A Physiologically Based Pharmacokinetic and Pharmacodynamic Model of Lixisenatide
Olivia Yau2, Matthias König1
1Humboldt-Universität zu Berlin, Institute for Theoretical Biology, Berlin, Germany, 2University of Toronto, Dept. of Pharmacology and Toxicology, Toronto, Canada
HIC conference 2025, Humboldt Internship Program Day, Berlin, 7 July 2025
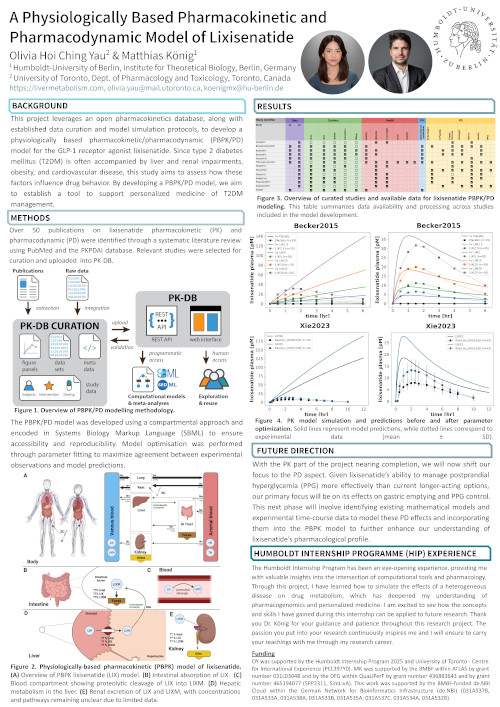
This project leverages an open pharmacokinetics database, along with established data curation and model simulation protocols, to develop a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model for the GLP-1 receptor agonist lixisenatide. Since type 2 diabetes mellitus (T2DM) is often accompanied by liver and renal impairments, obesity, and cardiovascular disease, this study aims to assess how these factors influence drug behavior. By developing a PBPK/PD model, we aim to establish a tool to support personalized medicine of T2DM management.
Keywords:
pharmacokinetics,
pharmacodynamics,
lixisenatide,
GLP-1 agonists,
|
|
| 2025-07-07 |
|
A Physiologically Based Pharmacokinetic Model of Dulaglutide
Kim Minjun2, Matthias König1
1Humboldt-Universität zu Berlin, Institute for Theoretical Biology, Berlin, Germany, 2National University of Singapore, Singapore
HIC conference 2025, Humboldt Internship Program Day, Berlin, 7 July 2025
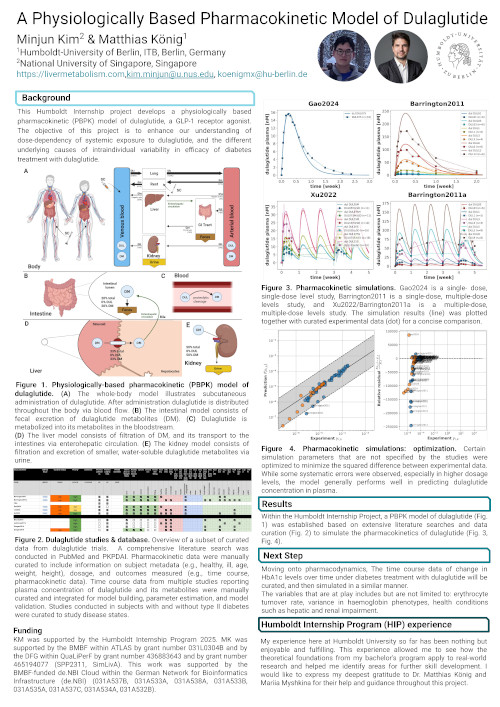
This Humboldt Internship project develops a physiologically based pharmacokinetic (PBPK) model of dulaglutide, a GLP-1 receptor agonist. The objective of this project is to enhance our understanding of dose-dependency of systemic exposure to dulaglutide, and the different underlying causes of intraindividual variability in efficacy of diabetes treatment with dulaglutide.
Keywords:
pharmacokinetics,
pharmacodynamics,
dulaglutide,
GLP-1 agonists,
|
|
| 2025-07-07 |
|
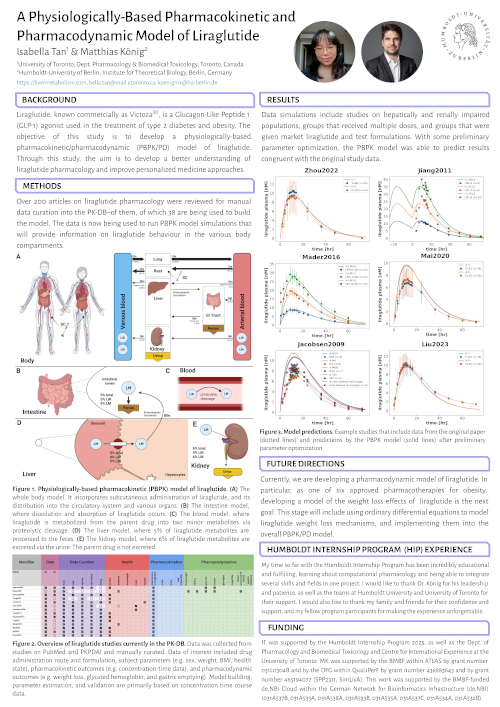
A Physiologically-Based Pharmacokinetic and Pharmacodynamic Model of Liraglutide
Isabella Tan2, Matthias König1
1Humboldt-Universität zu Berlin, Institute for Theoretical Biology, Berlin, Germany, 2University of Toronto, Dept. Pharmacology & Biomedical Toxicology, Toronto, Canada
HIC conference 2025, Humboldt Internship Program Day, Berlin, 7 July 2025
Liraglutide, known commercially as Victoza(R), is a Glucagon-Like Peptide 1 (GLP-1) agonist used in the treatment of type 2 diabetes and obesity. The objective of this study is to develop a physiologically-based pharmacokinetic/pharmacodynamic (PBPK/PD) model of liraglutide. Through this study, the aim is to develop a better understanding of liraglutide pharmacology and improve personalized medicine approaches.
Keywords:
pharmacokinetics,
pharmacodynamics,
liraglutide,
GLP-1 agonists,
|
|
| 2024-11-05 |
|
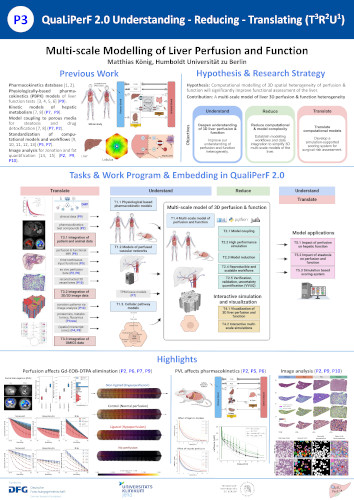
Multi-scale Modelling of Liver Perfusion and Function
Matthias König1
1Humboldt-Universität zu Berlin, Faculty of Life Science, Institute for Biology, Systems Medicine of the Liver, Berlin, Germany
Qualiperf 2 Defense, 5 November 2024
Hypothesis: Computational modelling of 3D spatial heterogeneity of perfusion & function will significantly improve functional assessment of the liver. Contribution: A multi-scale model of liver 3D perfusion & function heterogeneity.
Keywords:
pharmacokinetics,
pharmacodynamics,
digital twin,
liver,
perfusion,
function,
|
|
| 2024-11-05 |

|
Data Management Project
Matthias König1, Hans-Michael Tautenhahn2
1Humboldt-Universität zu Berlin, Faculty of Life Science, Institute for Biology, Systems Medicine of the Liver, Berlin, Germany 2Department of Visceral, Transplantation, Thoracic and Vascular Surgery, Universitätsklinikum Leipzig
Qualiperf 2 Defense, 5 November 2024
Central data management project.
Keywords:
data management,
FAIR,
|
|
| 2024-07-18 |

|
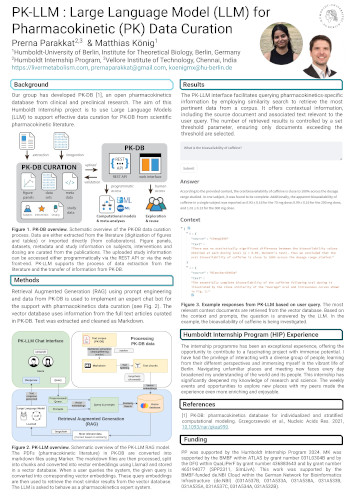
PK-LLM : Large Language Model (LLM) for Pharmacokinetic (PK) Data Curation
Prerna Parakkat2, Matthias König1
1Humboldt-University of Berlin, Institute for Theoretical Biology, Berlin, Germany, 2Vellore Institute of Technology, Chennai, India
HIC conference 2024, Humboldt Internship Program Day, Berlin, 18 July 2024
Our group has developed PK-DB, an open pharmacokinetics database from clinical and preclinical research. The aim of this Humboldt Internship project is to use Large Language Models (LLM) to support effective data curation for PK-DB from scientific pharmacokinetic literature.
Keywords:
LLM,
large language models,
pharmacometrics,
pharmacokinetics,
|
|
| 2024-07-18 |
|
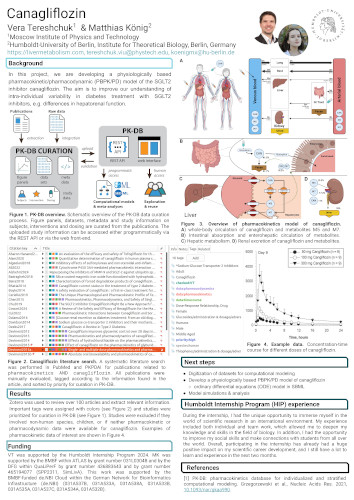
Canagliflozin
Vera Tereshchuk1, Matthias König2
1Moscow Institute of Physics and Technology, 2Humboldt-University of Berlin, Institute for Theoretical Biology, Berlin, Germany
HIC conference 2024, Humboldt Internship Program Day, Berlin, 18 July 2024
In this project, we are developing a physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of the SGLT2 inhibitor canagliflozin. The aim is to improve our understanding of intra-individual variability in diabetes treatment with SGLT2 inhibitors, e.g. differences in hepatorenal function.
Keywords:
canagliflozin,
pharmacometrics,
|
|
| 2024-07-18 |
|
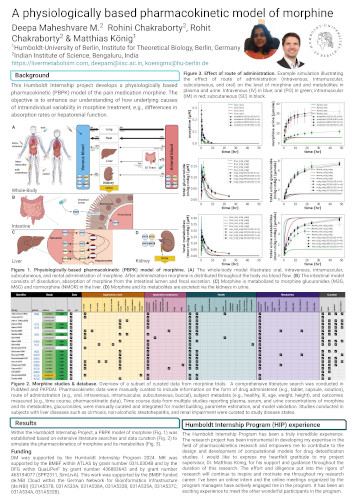
A physiologically based pharmacokinetic model of morphine
Deepa Maheshvare M.2, Rohini Chakraborty2, Rohit Chakraborty2, Matthias König1
2Humboldt-University of Berlin, Institute for Theoretical Biology, Berlin, Germany, 2Indian Institute of Science, Bengaluru, India
HIC conference 2024, Humboldt Internship Program Day, Berlin, 18 July 2024
This Humboldt Internship project develops a physiologically based pharmacokinetic (PBPK) model of the pain medication morphine. The objective is to enhance our understanding of how underlying causes of intraindividual variability in morphine treatment, e.g., differences in absorption rates or hepatorenal function.
Keywords:
morphine,
pharmacometrics,
|
|
| 2024-06-26 |
|
Captopril in Focus: Establishing an Open Pharmacokinetic Dataset and PBPK Modeling
Mariia Myshkina1, Matthias König1
1Humboldt-University of Berlin, Institute for Theoretical Biology, Berlin, Germany
PAGE2024 - Population Approach Group Europe, Rome, 26-28 June 2024
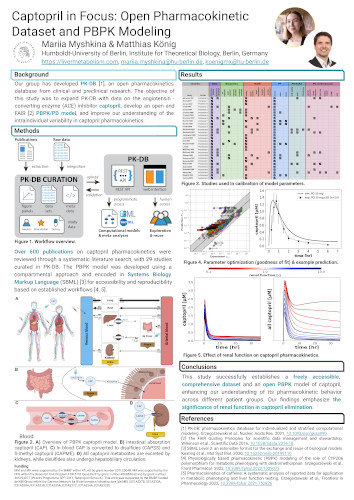
Introduction: Angiotensin-converting enzyme (ACE) inhibitors, such as captopril, are widely prescribed for the treatment of hypertension and heart failure. Captopril, the first ACE inhibitor developed in the 1980s, is primarily taken orally and is rapidly absorbed, reaching peak plasma concentrations within an hour. Its half-life is approximately 2 hours, but several factors, including renal function, heart disease, age and sex, can affect its pharmacokinetics. Despite the widespread use of captopril, comprehensive pharmacokinetic data have been lacking. Our group has developed PK-DB (https://pk-db.com) [1], an open database containing high-quality pharmacokinetic data from clinical and preclinical research. PK-DB includes patient cohort characteristics, interventions, concentration-time profiles, and kinetic parameters, with the ability to automatically calculate pharmacokinetic parameters using non-compartmental methods.
Objectives: The objectives of this study were to curate and analyze captopril pharmacokinetic data, expand the PK-DB, develop a whole-body physiologically based pharmacokinetic/pharmacodynamic (PBPK/PD) model of captopril in accordance with FAIR principles [2], and improve understanding of the intra-individual variability in captopril pharmacokinetics influenced by factors such as age, sex, renal and cardiac function.
Methods: Over 600 publications on captopril pharmacokinetics were reviewed through a systematic literature search, with a subset selected for data curation in PK-DB. The PBPK model was developed using a compartmental approach and encoded in Systems Biology Markup Language (SBML) [3] for accessibility and reproducibility.
Results: A comprehensive set of captopril pharmacokinetic data was generated from 15 clinical studies in healthy volunteers and patients with renal impairment, heart failure and hypertension. The PBPK model includes four major compartments (blood, intestine, liver, kidney) and describes absorption, distribution, metabolism and excretion. It successfully represents the pharmacokinetic profile of captopril in different age groups, showing minimal age-related differences in unchanged drug concentration. Heart failure did not significantly affect maximum plasma concentration or area under the curve (AUC), but renal function strongly influenced captopril elimination. We present this data set and open PBPK model of captopril as a valuable resource for further research.
Conclusions: This study successfully establishes a freely accessible, comprehensive dataset and an open PBPK model of captopril, enhancing our understanding of its pharmacokinetic behavior across different patient groups. Our findings emphasize the significance of renal function in captopril elimination and provide a valuable resource for personalized medicine approaches in hypertension and heart failure treatment.
References: [1] Grzegorzewski, J. et al. 2021. PK-DB: pharmacokinetics database for individualized and stratified computational modeling. Nucleic Acids Research. 49, D1 (Jan. 2021), D1358–D1364. DOI:https://doi.org/10.1093/nar/gkaa990. [2] Wilkinson, M.D. et al. 2016. The FAIR Guiding Principles for scientific data management and stewardship. Scientific Data. 3, (Mar. 2016), 160018. DOI:https://doi.org/10.1038/sdata.2016.18. [3] Keating, SM. et al, SBML Level 3: an extensible format for the exchange and reuse of biological models. Mol Syst Biol. 2020 Aug;16(8):e9110. doi: 10.15252/msb.20199110.
Keywords:
captopril,
pharmacometrics,
|
|
| 2010-06-01 |
|
Blood glucose control by the human hepatocyte
Matthias König1, Sascha Bulik1, Hermann-Georg Holzhütter1
1Charité University Medicine Berlin, Insitute of Biochemistry, Berlin, Germany
Spetses 2010
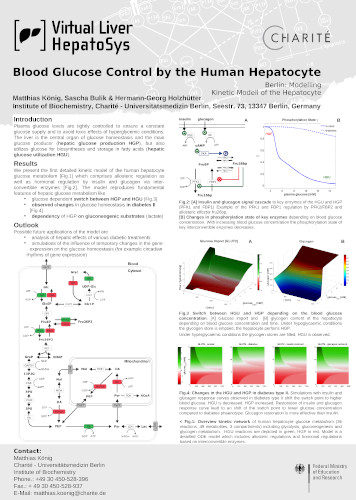
Introduction: Plasma glucose levels are tightly controlled to ensure a constant glucose supply and to avoid toxic effects of hyperglycemic conditions. The liver is the central organ of glucose homeostasis and the main glucose producer (hepatic glucose production HGP), but also utilizes glucose for biosyntheses and storage in fatty acids (hepatic glucose utilization HGU). Results: We present the first detailed kinetic model of the human hepatocyte glucose metabolism [Fig.1] which comprises allosteric regulation as well as hormonal regulation by insulin and glucagon via inter-convertible enzymes [Fig.2]. The model reproduces fundamental features of hepatic glucose metabolism like glucose dependent switch between HGP and HGU [Fig.3] observed changes in glucose homeostasis in diabetes II [Fig.4] dependency of HGP on gluconeogenic substrates (lactate). Outlook: Possible future applications of the model are analysis of hepatic effects of various diabetic treatments simulations of the influence of temporary changes in the gene expression on the glucose homeostasis (for example circadian rhythms of gene expression).
Keywords:
glucose,
modelling,
|
|